This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Journal:Cell:1
From Proteopedia
(Difference between revisions)

| Line 12: | Line 12: | ||
[[Image:IFNa_IFNAR2_interaction_map.png|300px||right|]] | [[Image:IFNa_IFNAR2_interaction_map.png|300px||right|]] | ||
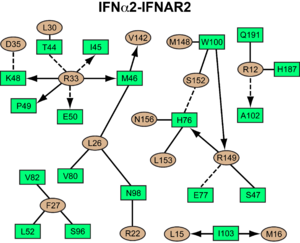
<scene name='User:David_Canner/Workbench/Opening_ifna/2'>Interferon</scene> interacts primarily with the <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction/1'>D1 domain of IFNAR2</scene>. Arg33(IFN) appears to be the <scene name='User:David_Canner/Workbench2/Ifn_arg_33/1'>single most important residue</scene> for the interaction of the IFN ligand with IFNAR2. It forms an <scene name='User:David_Canner/Workbench2/Ifn_h_bonds_cartoon/2'>extensive hydrogen-bonding network</scene> with the main chain carbonyl oxygen atoms of <scene name='User:David_Canner/Workbench2/Ifn_h_bonds_non_cartoon/3'>Ile45(IFNAR2) and Glu50(IFNAR2) and the side chain of Thr44(IFNAR2)</scene>. This residue is present in IFNa, IFNw, IFNb and IFNe. Two hydrophobic interaction clusters are part of the <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interact_hydro_full/1'>IFNa-IFNAR2</scene> interface: <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_hydrop1/3'>the first one</scene> is formed between Leu15 and Met16 of the IFN molecule and Trp100 and Ile103 of IFNAR2; <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_hydrop2/1'>the second one</scene> comprises Leu26, Phe27, Leu30 and Val142 of the ligand and Met46, Leu52, Val80 and the methyl group of Thr44 of the receptor. Replacing <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_leu_30/1'>Leu30(IFN) with alanine</scene> reduces affinity by three orders of magnitude (the second most important residue for binding). This is surprising, as it is <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_leu_30_nono/4'>not engaged in any intimate contacts with IFNAR2 residues</scene>. One reason for its importance might be a <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_arg_stabilized/1'>stabilizing effect on the position of Arg33(IFN)</scene>. | <scene name='User:David_Canner/Workbench/Opening_ifna/2'>Interferon</scene> interacts primarily with the <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction/1'>D1 domain of IFNAR2</scene>. Arg33(IFN) appears to be the <scene name='User:David_Canner/Workbench2/Ifn_arg_33/1'>single most important residue</scene> for the interaction of the IFN ligand with IFNAR2. It forms an <scene name='User:David_Canner/Workbench2/Ifn_h_bonds_cartoon/2'>extensive hydrogen-bonding network</scene> with the main chain carbonyl oxygen atoms of <scene name='User:David_Canner/Workbench2/Ifn_h_bonds_non_cartoon/3'>Ile45(IFNAR2) and Glu50(IFNAR2) and the side chain of Thr44(IFNAR2)</scene>. This residue is present in IFNa, IFNw, IFNb and IFNe. Two hydrophobic interaction clusters are part of the <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interact_hydro_full/1'>IFNa-IFNAR2</scene> interface: <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_hydrop1/3'>the first one</scene> is formed between Leu15 and Met16 of the IFN molecule and Trp100 and Ile103 of IFNAR2; <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_hydrop2/1'>the second one</scene> comprises Leu26, Phe27, Leu30 and Val142 of the ligand and Met46, Leu52, Val80 and the methyl group of Thr44 of the receptor. Replacing <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_leu_30/1'>Leu30(IFN) with alanine</scene> reduces affinity by three orders of magnitude (the second most important residue for binding). This is surprising, as it is <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_leu_30_nono/4'>not engaged in any intimate contacts with IFNAR2 residues</scene>. One reason for its importance might be a <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_arg_stabilized/1'>stabilizing effect on the position of Arg33(IFN)</scene>. | ||
| - | Most of the residues involved in the | + | Most of the residues involved in the IFNa2-IFNAR2 interaction are also found in the IFNw-IFNAR2 interface of the IFNw ternary complex. |
[[Image:IFNw_IFNAR2_interaction_map.png|300px|left|]] | [[Image:IFNw_IFNAR2_interaction_map.png|300px|left|]] | ||
A significant difference in the IFNAR2 interface between <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction_dif/5'>IFNa2</scene> and IFNw is related to <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction_salt/1'>Arg149 in IFNa2</scene>, which is replaced with Lys152 in <scene name='User:David_Canner/Workbench2/Ifnw_ifnar21_structure/3'>IFNw</scene>. In the <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_interface/3'>IFNw-IFNAR2 interface</scene>, this residue forms an <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_salt/1'>intramolecular salt bridge</scene> with Glu147(IFN), but <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_no_interact/1'>does not contact Glu77 of the receptor</scene>. The IFNa-IFNAR2 interface buries ~1841 Å^2 of surface area. | A significant difference in the IFNAR2 interface between <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction_dif/5'>IFNa2</scene> and IFNw is related to <scene name='User:David_Canner/Workbench2/Ifn_ifnar2_interaction_salt/1'>Arg149 in IFNa2</scene>, which is replaced with Lys152 in <scene name='User:David_Canner/Workbench2/Ifnw_ifnar21_structure/3'>IFNw</scene>. In the <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_interface/3'>IFNw-IFNAR2 interface</scene>, this residue forms an <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_salt/1'>intramolecular salt bridge</scene> with Glu147(IFN), but <scene name='User:David_Canner/Workbench2/Ifnw_ifnar2_no_interact/1'>does not contact Glu77 of the receptor</scene>. The IFNa-IFNAR2 interface buries ~1841 Å^2 of surface area. | ||
Revision as of 08:52, 25 July 2011
| |||||||||||
- ↑ no reference
Proteopedia Page Contributors and Editors (what is this?)
Christoph Thomas, Jaime Prilusky, Joel L. Sussman, Michal Harel, Alexander Berchansky
This page complements a publication in scientific journals and is one of the Proteopedia's Interactive 3D Complement pages. For aditional details please see I3DC.