This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Tom Sandbox
From Proteopedia
(Difference between revisions)
| Line 6: | Line 6: | ||
===GCN4 - Leucine Zipper=== | ===GCN4 - Leucine Zipper=== | ||
| - | + | Blah blah information about leucine zipper GCN4. | |
{{STRUCTURE_1ysa| PDB=1ysa | SCENE= }} | {{STRUCTURE_1ysa| PDB=1ysa | SCENE= }} | ||
==About this Structure== | ==About this Structure== | ||
| - | [[2zta]] is a | + | [[2zta]] is composed of two identical 52 residue alpha helix chains that grouped together to form a dimer. The dimer binds through interlocking leucine amino acids in the C terminal ends, while pinching in on the major groove of DNA in the N terminal end. |
==See Also== | ==See Also== | ||
| - | + | [[2zta]] | |
| + | [[1ysa]] | ||
==Reference== | ==Reference== | ||
| - | <ref group="xtra">PMID:1557122</ref><references group="xtra"/> | ||
| - | [[Category: Saccharomyces cerevisiae]] | ||
| - | [[Category: Carey, M.]] | ||
| - | [[Category: Harrison, S C.]] | ||
| - | [[Category: Marmorstein, R.]] | ||
| - | [[Category: Ptashne, M.]] | ||
| - | [[Category: Double helix]] | ||
| - | [[Category: Protein-dna complex]] | ||
| - | [[Category: Transcription/dna complex]] | ||
Revision as of 00:34, 9 November 2011
| |||||||
| 2zta, resolution 1.80Å () | |||||||
|---|---|---|---|---|---|---|---|
| Non-Standard Residues: | |||||||
| |||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
Contents |
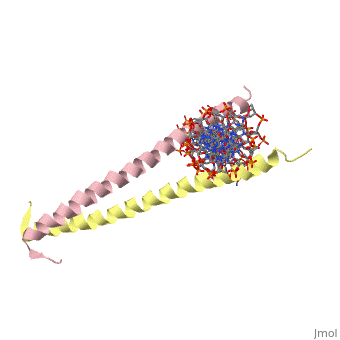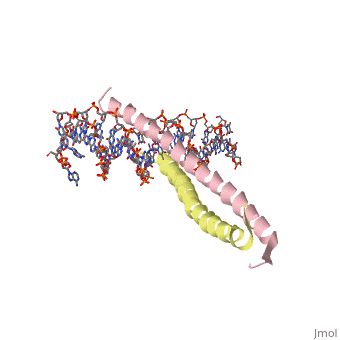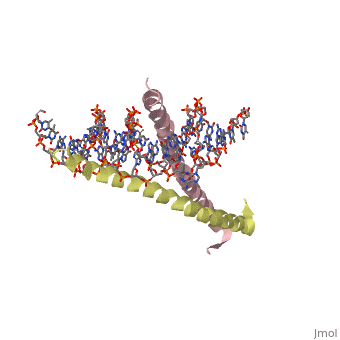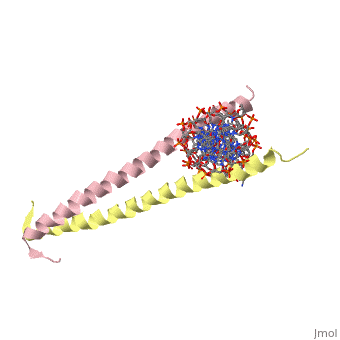
GCN4 - Leucine Zipper
Blah blah information about leucine zipper GCN4.
| |||||||||
| 1ysa, resolution 2.90Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
About this Structure
2zta is composed of two identical 52 residue alpha helix chains that grouped together to form a dimer. The dimer binds through interlocking leucine amino acids in the C terminal ends, while pinching in on the major groove of DNA in the N terminal end.