This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
A-ATP Synthase
From Proteopedia
| Line 6: | Line 6: | ||
<StructureSection load=1e79 size='500' side='right' caption='F1-ATP synthase motor domain', ([[1e79]])' scene=''> | <StructureSection load=1e79 size='500' side='right' caption='F1-ATP synthase motor domain', ([[1e79]])' scene=''> | ||
| - | == | + | ==<scene name='A-ATP_Synthase/Rotary_stalk/1'>F1 ATP Synthase Rotary Mechanism</scene>== |
| - | The central stalk in ATP synthase, made of gamma, delta and epsilon subunits in the mitochondrial enzyme, is the key rotary element in the enzyme's catalytic mechanism. The | + | The central stalk in ATP synthase, made of gamma, delta and epsilon subunits in the mitochondrial enzyme, is the key rotary element in the enzyme's catalytic mechanism. The <scene name='A-ATP_Synthase/Rotary_stalk/2'>Gamma subunit</scene> penetrates the catalytic (alpha beta)(3) domain and protrudes beneath it, interacting with a ring of c subunits in the membrane that drives rotation of the stalk during ATP synthesis. The delta and epsilon subunits interact with a Rossmann fold in the gamma subunit, forming a foot. In ATP synthase, this foot interacts with the c-ring and couples the transmembrane proton motive force to catalysis in the (<scene name='A-ATP_Synthase/Rotary_stalk/4'>alpha</scene> <scene name='A-ATP_Synthase/Rotary_stalk/3'>beta</scene>)(3) domain. |
When operating as a generator, it uses the power of rotational motion to build ATP, or when operating as a motor, it breaks down ATP to spin the axle the opposite direction. The synthesis of ATP requires several steps, including the binding of ADP and phosphate, the formation of the new phosphate-phosphate bond, and release of ATP. As the axle turns, it '''forces the motor into three different conformations that assist these difficult steps.''' | When operating as a generator, it uses the power of rotational motion to build ATP, or when operating as a motor, it breaks down ATP to spin the axle the opposite direction. The synthesis of ATP requires several steps, including the binding of ADP and phosphate, the formation of the new phosphate-phosphate bond, and release of ATP. As the axle turns, it '''forces the motor into three different conformations that assist these difficult steps.''' | ||
Revision as of 19:31, 28 November 2011
Structure
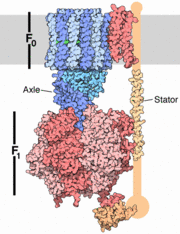
The structure on the right shows the F1 motor and the axle that connects the two. ATP synthesis is composed of two rotary motors, each powered by a different fuel. The motor at the top, termed F0, an electric motor. It is embedded in a membrane (shown schematically as a gray stripe here), and is powered by the flow of hydrogen ions across the membrane. As the protons flow through the motor, they turn a circular rotor . This rotor is connected to the second motor, termed F1. The F1 motor is a chemical motor, powered by ATP. The two motors are connected together by a stator, shown on the right, so that when F0 turns, F1 turns too. A-ATP synthase is very similar to F ATP Synthase and is composed of two parts A1 and A0 which are composed of at least nine subunits A3B3C:D:E:F:H2:a:cx that function as a pair of rotary motors connected by central and peripheral stalk(s) [1]. The A0 domain is the hydrophobic membrane embedded ion-translocating sector that uses the H+ gradient to power ATP synthase in domain A1. A1 is catalytic and water soluble containing A and B subunits. These subunits are comparable to F-ATP synthase ATP synthase alpha/beta subunits. The A subunit of A1 is catalytic and the B subunit is regulatory, with a substrate-binding site on each. [1]
| |||||||||||
| |||||||||||
| |||||||
| Transition state, 1e79 | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | , , , , , | ||||||
| Activity: | H(+)-transporting two-sector ATPase, with EC number 3.6.3.14 | ||||||
| Related: | 1bmf, 1cow, 1e1q, 1e1r, 1efr, 1nbm, 1qo1
| ||||||
| |||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
Contents |
References
- ↑ 1.0 1.1 Muller V, Lemker T, Lingl A, Weidner C, Coskun U, Gruber G. Bioenergetics of archaea: ATP synthesis under harsh environmental conditions. J Mol Microbiol Biotechnol. 2005;10(2-4):167-80. PMID:16645313 doi:10.1159/000091563
- ↑ Gibbons C, Montgomery MG, Leslie AG, Walker JE. The structure of the central stalk in bovine F(1)-ATPase at 2.4 A resolution. Nat Struct Biol. 2000 Nov;7(11):1055-61. PMID:11062563 doi:10.1038/80981
- ↑ Schafer IB, Bailer SM, Duser MG, Borsch M, Bernal RA, Stock D, Gruber G. Crystal structure of the archaeal A1Ao ATP synthase subunit B from Methanosarcina mazei Go1: Implications of nucleotide-binding differences in the major A1Ao subunits A and B. J Mol Biol. 2006 May 5;358(3):725-40. Epub 2006 Mar 10. PMID:16563431 doi:http://dx.doi.org/10.1016/j.jmb.2006.02.057
- ↑ Gonzalez JM, Masuchi Y, Robb FT, Ammerman JW, Maeder DL, Yanagibayashi M, Tamaoka J, Kato C. Pyrococcus horikoshii sp. nov., a hyperthermophilic archaeon isolated from a hydrothermal vent at the Okinawa Trough. Extremophiles. 1998 May;2(2):123-30. PMID:9672687
- ↑ Manimekalai MS, Kumar A, Jeyakanthan J, Gruber G. The Transition-Like State and P(i) Entrance into the Catalytic A Subunit of the Biological Engine A-ATP Synthase. J Mol Biol. 2011 Mar 16. PMID:21396943 doi:10.1016/j.jmb.2011.03.010
- ↑ Priya R, Kumar A, Manimekalai MS, Gruber G. Conserved Glycine Residues in the P-Loop of ATP Synthases Form a Doorframe for Nucleotide Entrance. J Mol Biol. 2011 Sep 8. PMID:21925186 doi:10.1016/j.jmb.2011.08.045
- ↑ Manimekalai MS, Kumar A, Jeyakanthan J, Gruber G. The Transition-Like State and P(i) Entrance into the Catalytic A Subunit of the Biological Engine A-ATP Synthase. J Mol Biol. 2011 Mar 16. PMID:21396943 doi:10.1016/j.jmb.2011.03.010
Proteopedia Page Contributors and Editors (what is this?)
Kaitlin Chase MacCulloch, Michal Harel, Alexander Berchansky