This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
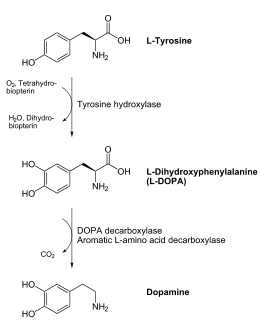
DOPA decarboxylase
From Proteopedia
(Difference between revisions)
| Line 37: | Line 37: | ||
---- | ---- | ||
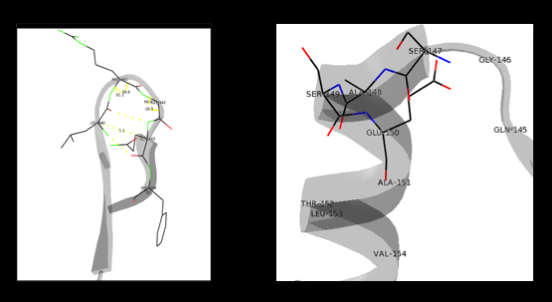
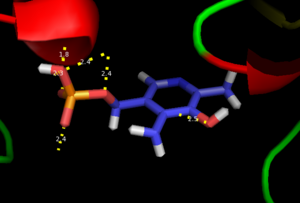
As well, a '''salt bridge''' exists between the carboxyl group of <scene name='DOPA_decarboxylase/Aspartic/1'>Asp271</scene> and the protonated pyridine nitrogen of PLP to further stabilize intermediate. Essentially, a salt bridge combines hydrogen bonding and electrostatic interactions (two common types non-covalent interactions). This interaction serves to provide an electron sink that can stabilize the carbanionic intermediates <ref name="jansonius">PMID:9914259 </ref> . PLP is further anchored to the protein by an extended '''hydrogen bond network''', as shown below. | As well, a '''salt bridge''' exists between the carboxyl group of <scene name='DOPA_decarboxylase/Aspartic/1'>Asp271</scene> and the protonated pyridine nitrogen of PLP to further stabilize intermediate. Essentially, a salt bridge combines hydrogen bonding and electrostatic interactions (two common types non-covalent interactions). This interaction serves to provide an electron sink that can stabilize the carbanionic intermediates <ref name="jansonius">PMID:9914259 </ref> . PLP is further anchored to the protein by an extended '''hydrogen bond network''', as shown below. | ||
| - | [[image:h-bonding.png|thumb|center| | + | [[image:h-bonding.png|thumb|center|300px|'''H-bonding network of PLP in the active site''']] |
---- | ---- | ||
The only two active site residues from the adjacent monomer, Ile-101 and Phe-103, are part of the substrate binding pocket. | The only two active site residues from the adjacent monomer, Ile-101 and Phe-103, are part of the substrate binding pocket. | ||
| Line 43: | Line 43: | ||
===Inhibitor Binding=== | ===Inhibitor Binding=== | ||
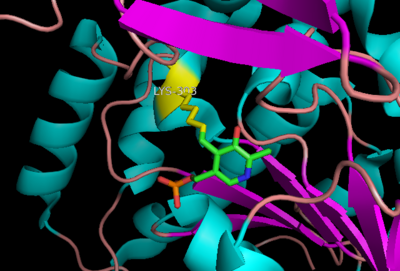
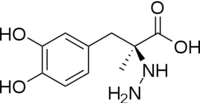
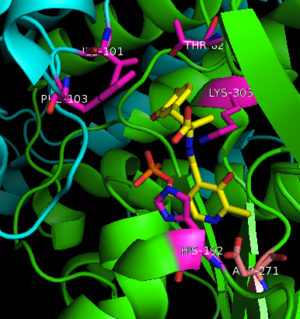
[[image:carbiDOPA.png|thumb|left|200px|'''carbiDOPA''']]The inhibitor <scene name='DOPA_decarboxylase/Carbidopa/1'>carbiDOPA</scene> binds to the enzyme by forming a hydrazone linkage with PLP through its hydrazine moiety. The catechol ring of carbiDOPA is deeply buried in the active site cleft and is stabilized by <scene name='DOPA_decarboxylase/Vanderwaals/1'>van der waals contact</scene> with Ile-101 and Phe-103. The 4' hydroxyl group of the catechol ring participates in hydrogen bonding with <scene name='DOPA_decarboxylase/Thr-82/1'>Thr-82</scene>, further stabilizing the inhibitor in the active site cleft. PLP is further involved in substrate binding by forming a hydrogen bond to the 3' of the catechol ring. <scene name='DOPA_decarboxylase/His192/1'>His-192</scene>, a highly conserved residue of PLP-dependent decarboxylases <ref name="ishii">PMID:8889823 </ref> hydrogen bonds to the carboxylate group of carbiDOPA. | [[image:carbiDOPA.png|thumb|left|200px|'''carbiDOPA''']]The inhibitor <scene name='DOPA_decarboxylase/Carbidopa/1'>carbiDOPA</scene> binds to the enzyme by forming a hydrazone linkage with PLP through its hydrazine moiety. The catechol ring of carbiDOPA is deeply buried in the active site cleft and is stabilized by <scene name='DOPA_decarboxylase/Vanderwaals/1'>van der waals contact</scene> with Ile-101 and Phe-103. The 4' hydroxyl group of the catechol ring participates in hydrogen bonding with <scene name='DOPA_decarboxylase/Thr-82/1'>Thr-82</scene>, further stabilizing the inhibitor in the active site cleft. PLP is further involved in substrate binding by forming a hydrogen bond to the 3' of the catechol ring. <scene name='DOPA_decarboxylase/His192/1'>His-192</scene>, a highly conserved residue of PLP-dependent decarboxylases <ref name="ishii">PMID:8889823 </ref> hydrogen bonds to the carboxylate group of carbiDOPA. | ||
| - | [[image:actsite.png|thumb|center| | + | [[image:actsite.png|thumb|center|300px|'''Key interactions between the active site residues, PLP, and carbiDOPA''']] |
---- | ---- | ||
===Flexible Loop=== | ===Flexible Loop=== | ||
Revision as of 01:44, 20 January 2012
| |||||||||||
3D structures of DOPA decarboxylase
Update November 2011
3k40 – DDC – Drosophila melanogaster
1js3 – pDDC + inhibitor – pig
1js6 - pDDC
3rbf, 3rbl – hDDC – human
3rch – hDDC + vitamin B6 phosphate + pyridoxal phosphate
References
- ↑ 1.0 1.1 Schneider G, Kack H, Lindqvist Y. The manifold of vitamin B6 dependent enzymes. Structure. 2000 Jan 15;8(1):R1-6. PMID:10673430
- ↑ Miles EW. The tryptophan synthase alpha 2 beta 2 complex. Cleavage of a flexible loop in the alpha subunit alters allosteric properties. J Biol Chem. 1991 Jun 15;266(17):10715-8. PMID:1904055
- ↑ Burkhard P, Dominici P, Borri-Voltattorni C, Jansonius JN, Malashkevich VN. Structural insight into Parkinson's disease treatment from drug-inhibited DOPA decarboxylase. Nat Struct Biol. 2001 Nov;8(11):963-7. PMID:11685243 doi:http://dx.doi.org/10.1038/nsb1101-963
- ↑ Percudani R, Peracchi A. A genomic overview of pyridoxal-phosphate-dependent enzymes. EMBO Rep. 2003 Sep;4(9):850-4. PMID:12949584 doi:http://dx.doi.org/10.1038/sj.embor.embor914
- ↑ Aurora R, Rose GD. Helix capping. Protein Sci. 1998 Jan;7(1):21-38. PMID:9514257 doi:10.1002/pro.5560070103
- ↑ Jansonius JN. Structure, evolution and action of vitamin B6-dependent enzymes. Curr Opin Struct Biol. 1998 Dec;8(6):759-69. PMID:9914259
- ↑ 7.0 7.1 Ishii S, Mizuguchi H, Nishino J, Hayashi H, Kagamiyama H. Functionally important residues of aromatic L-amino acid decarboxylase probed by sequence alignment and site-directed mutagenesis. J Biochem. 1996 Aug;120(2):369-76. PMID:8889823
Proteopedia Page Contributors and Editors (what is this?)
Brittany Todd, David Canner, Michal Harel, Alexander Berchansky, Brian Hernandez