NADH quinone oxidoreductase (NQO1) with inhibitor dicoumarol
From Proteopedia
| Line 29: | Line 29: | ||
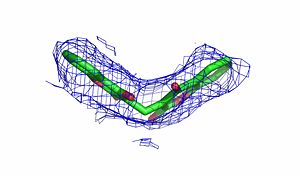
The quinone ES936 causes irreversible inhibition of the NQO1. <scene name='2f1o/Align2/5'>Alignment</scene> of the hNQO1 dimer in complex with <font color='red'><b>ES936 (red)</b></font> ([[1kbq]]) with the hNQO1−dicoumarol complex ([[2f1o]]) yields 0.45Å RMSD for the 546 Cα atoms. The ES936 causes structural change only in the position of Phe 232. The movement of this residue is smaller than that caused by dicoumarol. The <scene name='2f1o/Align2/6'>distance</scene> between Tyr 128 and Phe 232 in the hNQO1−ES936 complex is only ~7 Å, while in the hNQO1−dicoumarol complex it is ~12 Å. | The quinone ES936 causes irreversible inhibition of the NQO1. <scene name='2f1o/Align2/5'>Alignment</scene> of the hNQO1 dimer in complex with <font color='red'><b>ES936 (red)</b></font> ([[1kbq]]) with the hNQO1−dicoumarol complex ([[2f1o]]) yields 0.45Å RMSD for the 546 Cα atoms. The ES936 causes structural change only in the position of Phe 232. The movement of this residue is smaller than that caused by dicoumarol. The <scene name='2f1o/Align2/6'>distance</scene> between Tyr 128 and Phe 232 in the hNQO1−ES936 complex is only ~7 Å, while in the hNQO1−dicoumarol complex it is ~12 Å. | ||
</StructureSection> | </StructureSection> | ||
| + | |||
| + | ==3D structures of NQ01== | ||
| + | |||
| + | [[Quinone reductase]] | ||
Revision as of 09:54, 31 January 2012
Contents |
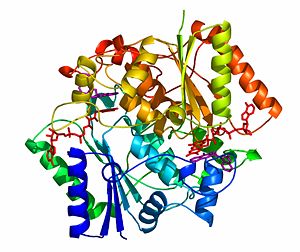
The crystal structure of NADH quinone oxidoreductase (NQO1) in complex with its potent inhibitor dicoumarol
NAD(P)H quinone oxidoreductase 1 (NQO1, EC 1.6.5.2) is a ubiquitous flavoenzyme that catalyzes two electron reduction of quinones to hydroquinones utilizing NAD(P)H as an electron donor. NQO1 is a homo-dimer that functions via a “ping pong” mechanism. NAD(P)H binds to NQO1, reduces the FAD co-factor and is then released, allowing the quinone substrate to bind the enzyme and to be reduced. The NAD(P)H and the quinone binding sites of NQO1 have a significant overlap, thus providing a molecular basis for this “ping pong” mechanism.
Certain coumarins, flavones and the reactive dye cibacron blue are competitive inhibitors of NQO1 activity, which compete with NAD(P)H for binding to NQO1. Dicoumarol (3-3’–methylene-bis (4-hydroxycoumarin)), is the most potent competitive inhibitor of NQO1. Dicoumarol competes with NAD(P)H for binding to NQO1 and prevents the electron transfer to FAD.In addition to its role in the detoxification of quinones, NQO1 is also a 20S proteasome-associated protein that plays an important role in the stability of the tumor suppressor p53 and several other short-lived proteins including p73α and ornithine decarboxylase (ODC, i.e. 7odc). NQO1 binds and stabilizes p53, protecting p53 from ubiquitin-independent 20S proteasomal degradation. Dicoumarol and several other inhibitors of NQO1 activity, which compete with NADH for binding to NQO1, disrupt the binding of NQO1 to p53 and induce ubiquitin-independent p53 degradation.
| |||||||||||
3D structures of NQ01
Additional Resources
For additional information, see: Carbohydrate Metabolism
Reference
The crystal structure of NAD(P)H quinone oxidoreductase 1 in complex with its potent inhibitor dicoumarol., Asher G, Dym O, Tsvetkov P, Adler J, Shaul Y, Biochemistry. 2006 May 23;45(20):6372-8. PMID:16700548
Proteopedia Page Contributors and Editors (what is this?)
Alexander Berchansky, Orly Dym, Michal Harel, Jaime Prilusky, Moshe Ben-David, Joel L. Sussman, David Canner, Eric Martz