We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
1mkf
From Proteopedia
(Difference between revisions)
OCA (Talk | contribs)
(New page: 200px <!-- The line below this paragraph, containing "STRUCTURE_1mkf", creates the "Structure Box" on the page. You may change the PDB parameter (which sets the PD...)
Next diff →
Revision as of 13:14, 5 June 2012
| |||||||
| 1mkf, resolution 2.10Å () | |||||||
|---|---|---|---|---|---|---|---|
| Gene: | M3 (Murid herpesvirus 4) | ||||||
| Related: | 1dok, 1ml0 | ||||||
| |||||||
| |||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB, TOPSAN | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
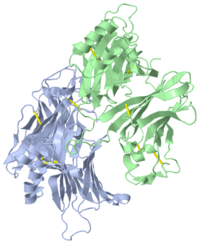
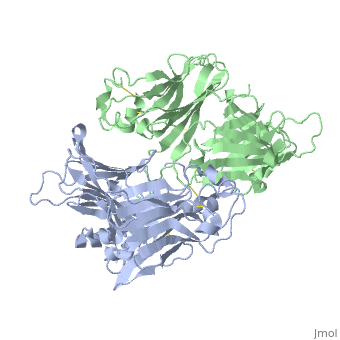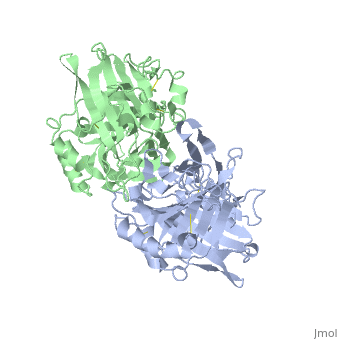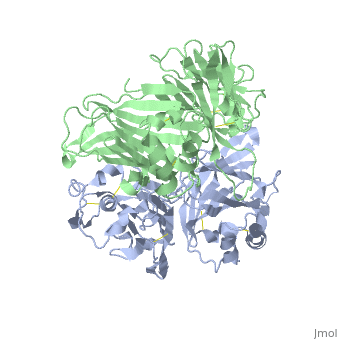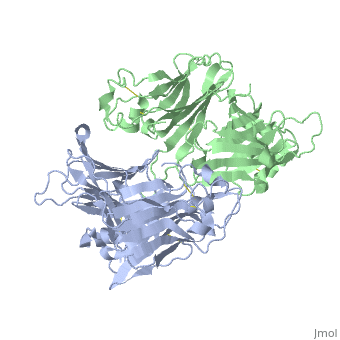
VIRAL CHEMOKINE BINDING PROTEIN M3 FROM MURINE GAMMAHERPESVIRUS 68
Template:ABSTRACT PUBMED 12419245
About this Structure
1mkf is a 2 chain structure with sequence from Murid herpesvirus 4. Full crystallographic information is available from OCA.
Reference
- Alexander JM, Nelson CA, van Berkel V, Lau EK, Studts JM, Brett TJ, Speck SH, Handel TM, Virgin HW, Fremont DH. Structural basis of chemokine sequestration by a herpesvirus decoy receptor. Cell. 2002 Nov 1;111(3):343-56. PMID:12419245
Categories: Murid herpesvirus 4 | Alexander, J M. | Fremont, D H. | MCSG, Midwest Center for Structural Genomics. | Chemokine binding protein | Decoy receptor | Herpesvirus | Immune system | Mcsg | Midwest center for structural genomic | Protein structure initiative | Psi | Structural genomic | Viral immune evasion