This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1mkf
From Proteopedia
(New page: 200px <!-- The line below this paragraph, containing "STRUCTURE_1mkf", creates the "Structure Box" on the page. You may change the PDB parameter (which sets the PD...) |
m (Protected "1mkf" [edit=sysop:move=sysop]) |
Revision as of 13:14, 5 June 2012
| |||||||
| 1mkf, resolution 2.10Å () | |||||||
|---|---|---|---|---|---|---|---|
| Gene: | M3 (Murid herpesvirus 4) | ||||||
| Related: | 1dok, 1ml0 | ||||||
| |||||||
| |||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB, TOPSAN | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
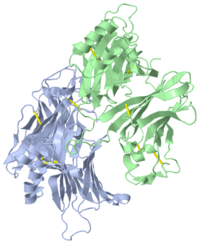
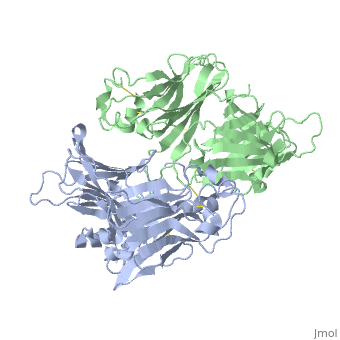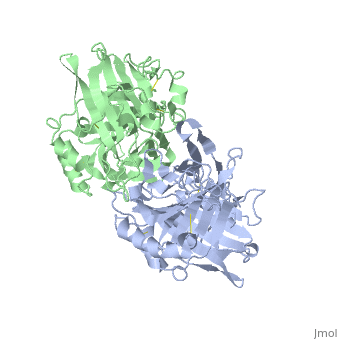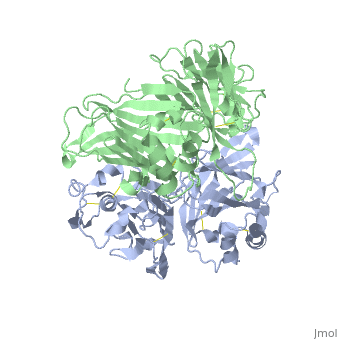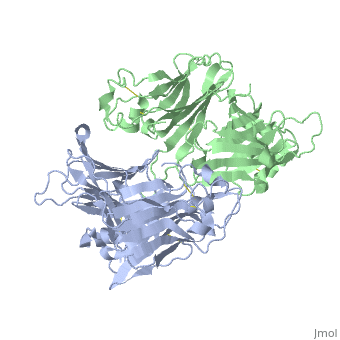
VIRAL CHEMOKINE BINDING PROTEIN M3 FROM MURINE GAMMAHERPESVIRUS 68
Template:ABSTRACT PUBMED 12419245
About this Structure
1mkf is a 2 chain structure with sequence from Murid herpesvirus 4. Full crystallographic information is available from OCA.
Reference
- Alexander JM, Nelson CA, van Berkel V, Lau EK, Studts JM, Brett TJ, Speck SH, Handel TM, Virgin HW, Fremont DH. Structural basis of chemokine sequestration by a herpesvirus decoy receptor. Cell. 2002 Nov 1;111(3):343-56. PMID:12419245
Categories: Murid herpesvirus 4 | Alexander, J M. | Fremont, D H. | MCSG, Midwest Center for Structural Genomics. | Chemokine binding protein | Decoy receptor | Herpesvirus | Immune system | Mcsg | Midwest center for structural genomic | Protein structure initiative | Psi | Structural genomic | Viral immune evasion