This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Protein SO0799
From Proteopedia
Alexander Berchansky (Talk | contribs)
(New page: 200px <!-- The line below this paragraph, containing "STRUCTURE_1yud", creates the "Structure Box" on the page. You may change the PDB parameter (which sets the PD...)
Next diff →
Revision as of 10:13, 6 June 2012
Contents |
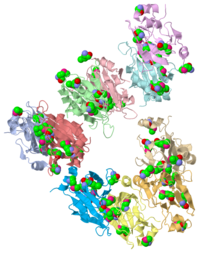
X-ray Crystal Structure of Protein SO0799 from Shewanella oneidensis. Northeast Structural Genomics Consortium Target SoR12.
About this Structure
1YUD is a 10 chains structure of sequences from Bacteria. Full crystallographic information is available from OCA.
Structural Informations
1yud chain a is member of the cupin superfamily, which includes proteins with a barrel-like structure. This superfamily includes 20 families and is one of the functionally most diverse known thus far, anyway, the functional site of members of this superfamily is generally located at the center of a conserved barrel. [1] The nine beta-strands of chain A are arranged in a beta-barrel-like structure, at the center of which there is a conserved domain ending in a heavy electronegative cavity, suggesting an interaction with positively charged molecules. The comparison of most structurally related proteins showed a conserved cluster specific for bivalent cations. This cluster was discovered analyzing data from Dali algorithm [2] and was confirmed by data from Skan algorithm [3]. The involved amino acids in this interactions are His55, Leu57, Glu61 and probably Mse111. His and Glu are almost always present in the aligned sequences. Leu and Mse represent two exceptions. There are no correspondences in the Dali-alignment with Leu and Mse, but, in this set of proteins, 1zvf chain a is the only one which binds a bivalent cation (Ni++) and that brings a sulfur-containing amino acid at the same time (Cys126). This can suggest that Ni++ is the interacting cation. Anyway, the most present cations in the annotated aligned sequences were Zn++ and Mn++, so one of these ions could be the one which actually binds the protein as well. These informations were obtained using tools provided by MarkUs [4] server.
References
- ↑ Garima Agarwal et al. 2009. Structure-Based Phylogeny as a Diagnostic for Functional Characterization of proteins with a Cupin Fold. PloS ONE. Volume 4.
- ↑ Holm L, Rosenström P. 2010. Dali server: conservation mapping in 3D. Nucl. Acids Res. 38, W545-549.
- ↑ Petrey D, Honig B. Methods Enzymol 2003, 374:492-509
- ↑ Markus Fischer. 2011. MarkUs: a server to navigate sequence–structure–function space. Oxford Journals. Life Sciences. Nucleic Acids Research. Volume39, Issuesuppl 2. Pp. W357-W361
Created with the participation of Francesco Aggazio.
Categories: Bacteria | Acton, T. | Chen, Y. | Forouhar, F. | Hunt, J F. | Kuzin, A P. | Ma, L C. | Montelione, G T. | NESG, Northeast Structural Genomics Consortium. | Tong, L. | Vorobiev, S. | Xiao, R. | Nesg | Northeast structural genomics consortium | Protein structure initiative | Psi | Q8e1n8 | Sor12 | X-ray structure