This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1one
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | {{Seed}} | ||
[[Image:1one.png|left|200px]] | [[Image:1one.png|left|200px]] | ||
| - | <!-- | ||
| - | The line below this paragraph, containing "STRUCTURE_1one", creates the "Structure Box" on the page. | ||
| - | You may change the PDB parameter (which sets the PDB file loaded into the applet) | ||
| - | or the SCENE parameter (which sets the initial scene displayed when the page is loaded), | ||
| - | or leave the SCENE parameter empty for the default display. | ||
| - | --> | ||
{{STRUCTURE_1one| PDB=1one | SCENE= }} | {{STRUCTURE_1one| PDB=1one | SCENE= }} | ||
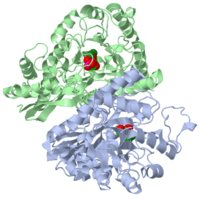
===YEAST ENOLASE COMPLEXED WITH AN EQUILIBRIUM MIXTURE OF 2'-PHOSPHOGLYCEATE AND PHOSPHOENOLPYRUVATE=== | ===YEAST ENOLASE COMPLEXED WITH AN EQUILIBRIUM MIXTURE OF 2'-PHOSPHOGLYCEATE AND PHOSPHOENOLPYRUVATE=== | ||
| - | |||
| - | <!-- | ||
| - | The line below this paragraph, {{ABSTRACT_PUBMED_8605183}}, adds the Publication Abstract to the page | ||
| - | (as it appears on PubMed at http://www.pubmed.gov), where 8605183 is the PubMed ID number. | ||
| - | --> | ||
{{ABSTRACT_PUBMED_8605183}} | {{ABSTRACT_PUBMED_8605183}} | ||
==About this Structure== | ==About this Structure== | ||
| - | + | [[1one]] is a 2 chain structure of [[Enolase]] with sequence from [http://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1ONE OCA]. | |
| + | |||
| + | ==See Also== | ||
| + | *[[Enolase|Enolase]] | ||
==Reference== | ==Reference== | ||
| - | <ref group="xtra">PMID: | + | <ref group="xtra">PMID:008605183</ref><references group="xtra"/> |
[[Category: Phosphopyruvate hydratase]] | [[Category: Phosphopyruvate hydratase]] | ||
[[Category: Saccharomyces cerevisiae]] | [[Category: Saccharomyces cerevisiae]] | ||
| Line 32: | Line 23: | ||
[[Category: Glycolysis]] | [[Category: Glycolysis]] | ||
[[Category: Lyase]] | [[Category: Lyase]] | ||
| - | |||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Mon Feb 16 17:22:38 2009'' | ||
Revision as of 18:54, 25 July 2012
| |||||||||
| 1one, resolution 1.80Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Non-Standard Residues: | |||||||||
| Activity: | Phosphopyruvate hydratase, with EC number 4.2.1.11 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
YEAST ENOLASE COMPLEXED WITH AN EQUILIBRIUM MIXTURE OF 2'-PHOSPHOGLYCEATE AND PHOSPHOENOLPYRUVATE
Template:ABSTRACT PUBMED 8605183
About this Structure
1one is a 2 chain structure of Enolase with sequence from Saccharomyces cerevisiae. Full crystallographic information is available from OCA.
See Also
Reference
- Larsen TM, Wedekind JE, Rayment I, Reed GH. A carboxylate oxygen of the substrate bridges the magnesium ions at the active site of enolase: structure of the yeast enzyme complexed with the equilibrium mixture of 2-phosphoglycerate and phosphoenolpyruvate at 1.8 A resolution. Biochemistry. 1996 Apr 9;35(14):4349-58. PMID:8605183 doi:http://dx.doi.org/10.1021/bi952859c