This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
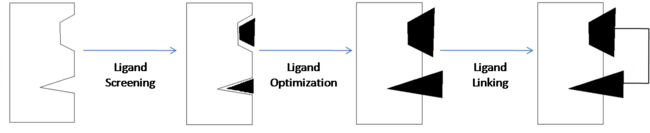
Fragment-Based Drug Discovery
From Proteopedia
(Difference between revisions)
| Line 43: | Line 43: | ||
Compounds 1 & 2 exhibited very poor binding affinity for Bcl-xl. The optimization of these two compounds resulted in <scene name='Sandbox_reserved_394/Compound_2/1'>Compound 3</scene>. In order to improve the binding affinity, the carboxylic acid of compound 1 was substituted with an acyl sulfonamide to capitalize on the hydrophilic interaction with the protein. This <scene name='Sandbox_reserved_394/Compound_2/3'>acylsulfonamide portion forms a hydrogen bond with Gly 142</scene> thereby increasing the affinity for Bcl-xl. The substitution of the sulfonamide actually allows the acidic proton to get closer to Gly 142 than it could in the carboxylic acid, which is why it is able to bind stronger to the amino acid. | Compounds 1 & 2 exhibited very poor binding affinity for Bcl-xl. The optimization of these two compounds resulted in <scene name='Sandbox_reserved_394/Compound_2/1'>Compound 3</scene>. In order to improve the binding affinity, the carboxylic acid of compound 1 was substituted with an acyl sulfonamide to capitalize on the hydrophilic interaction with the protein. This <scene name='Sandbox_reserved_394/Compound_2/3'>acylsulfonamide portion forms a hydrogen bond with Gly 142</scene> thereby increasing the affinity for Bcl-xl. The substitution of the sulfonamide actually allows the acidic proton to get closer to Gly 142 than it could in the carboxylic acid, which is why it is able to bind stronger to the amino acid. | ||
| - | Compound 2 was important in identifying the hydrophobicity of binding site 2 but was substituted with a <scene name='Sandbox_reserved_394/Nitro_thio_phenyl_sub/1'>3-nitro-4-(2-phenylthioethyl)aminophenyl group</scene>. This substitution more efficiently binds to site 2 through <scene name='Sandbox_reserved_394/Pi_stacking/ | + | Compound 2 was important in identifying the hydrophobicity of binding site 2 but was substituted with a <scene name='Sandbox_reserved_394/Nitro_thio_phenyl_sub/1'>3-nitro-4-(2-phenylthioethyl)aminophenyl group</scene>. This substitution more efficiently binds to site 2 through <scene name='Sandbox_reserved_394/Pi_stacking/3'>pi-pi interactions with Phe 101 and Tyr 199</scene>. This idea of using a known ligand to develop another ligand, and eventually drugs, is known as ligand-based drug design. |
{| class="wikitable collapsible collapsed" | {| class="wikitable collapsible collapsed" | ||
Revision as of 19:10, 7 November 2012
Drug Design: Fragment-Based Drug Discovery
| |||||||||||
References
- ↑ 1.0 1.1 Shuker S. B., Hajduk P. J., Meadows R. P., Fesik S. W. Discovering High-Affinity Ligands for Proteins: SAR by NMR. Science; Nov 29, 1996; 274, 5292; ProQuest Central pg. 1531.
- ↑ Oltersdorf T., Elmore S. W., Shoemaker A. R. An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Vol 435|2 June 2005|doi:10.1038/nature03579
- ↑ Pandit D. LIGAND-BASED DRUG DESIGN: I. CONFORMATIONAL STUDIES OF GBR 12909 ANALOGS AS COCAINE ANTAGONISTS; II. 3D-QSAR STUDIES OF SALVINORIN A ANALOGS AS εΑΡΡΑ OPIOID AGONISTS. http://archives.njit.edu/vol01/etd/2000s/2007/njit-etd2007-051/njit-etd2007-051.pdf