This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
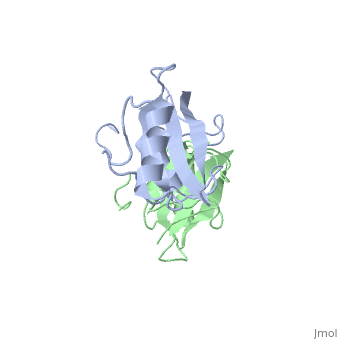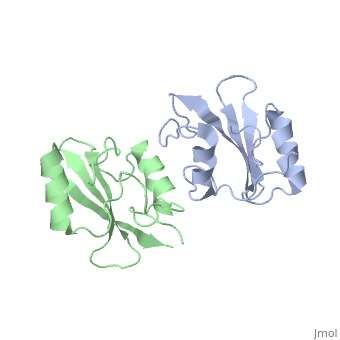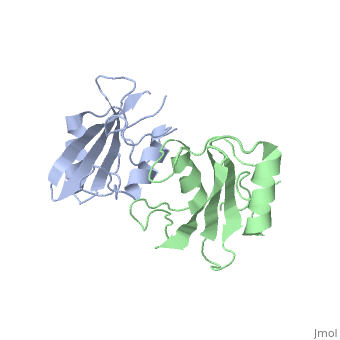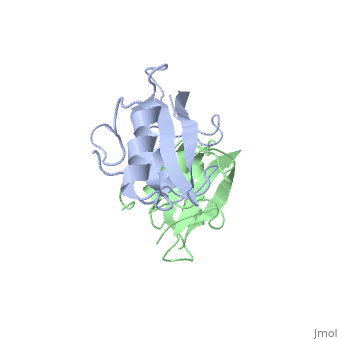
Grb10 SH2 Domain
From Proteopedia
(Difference between revisions)
| Line 18: | Line 18: | ||
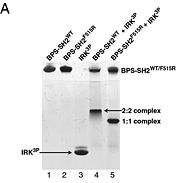
# BPS-SH2_WT (Wild Type) | # BPS-SH2_WT (Wild Type) | ||
# BPS-SH2_F515R (mutant = Phe515 --> Arg) | # BPS-SH2_F515R (mutant = Phe515 --> Arg) | ||
| - | # IRK_3P | + | # IRK_3P (tris-phosphorylated Insulin Receptor Kinase domain) |
# BPS-SH2_WT + IRK_3P | # BPS-SH2_WT + IRK_3P | ||
# BPS-SH2_F515R + IRK_3P | # BPS-SH2_F515R + IRK_3P | ||
As seen in lanes 1 and 2, the BPS-SH2 proteins did not travel down the gel due to their high pI; to resolve this issue, the researchers added IRK_3P to the two BPS-SH2 proteins which then made a complex that was mobile. <ref name=Guan>PMID: 12551896 </ref> Lane 4 shows a band labeled ''2:2 complex'' that shows the position of the SH2 dimer. The additional band found at the very top of lane 4 represents the BPS-SH2_WT protein that did not complex with high motility protein IRK_3P, i.e. it was not able to migrate through the gel due to its high pI. Lane 5 shows a band labeled ''1:1 complex'' elucidating that the Arg substitution at Phe515 did indeed produce a monomer, which was able to travel farther down the gel. | As seen in lanes 1 and 2, the BPS-SH2 proteins did not travel down the gel due to their high pI; to resolve this issue, the researchers added IRK_3P to the two BPS-SH2 proteins which then made a complex that was mobile. <ref name=Guan>PMID: 12551896 </ref> Lane 4 shows a band labeled ''2:2 complex'' that shows the position of the SH2 dimer. The additional band found at the very top of lane 4 represents the BPS-SH2_WT protein that did not complex with high motility protein IRK_3P, i.e. it was not able to migrate through the gel due to its high pI. Lane 5 shows a band labeled ''1:1 complex'' elucidating that the Arg substitution at Phe515 did indeed produce a monomer, which was able to travel farther down the gel. | ||
| + | |||
| + | '''Why BPS-SH2?''' | ||
| + | |||
| + | In order for the full-length Grb10 protein to interact with Insulin Receptor Kinase (IRK), the SH2 and BPS domain must be present. <ref name=Guan>PMID: 9506989 </ref> | ||
</StructureSection> | </StructureSection> | ||
Revision as of 07:26, 8 November 2012
| |||||||||||
Interaction Between Grb10 and E3 Ubiquitin Ligase NEDD4
WOOOOOOOO
|