This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Transfer RNA (tRNA)
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| + | <StructureSection load='' size='450' side='right' scene='1ehz/1ehz_default/3' caption=''> | ||
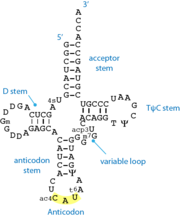
[[Image:TRNA.png|right|thumb|Standard 2D cloverleaf structure of tRNA. The shown example is methionine-specific tRNA from ''E.coli'' ]] | [[Image:TRNA.png|right|thumb|Standard 2D cloverleaf structure of tRNA. The shown example is methionine-specific tRNA from ''E.coli'' ]] | ||
'''tRNA''' or '''transfer RNA''' is stable, structured RNA present in all living cells. tRNA participates in the process of protein [[translation]] by the [[ribosome]]. Varying tRNA molecules carry a specific amino acid esterified on their 3'-OH group (the acceptor end). They also carry a specific triplet sequence, the '''anticodon''', which pairs with its complementary '''codon''' on the messenger RNA, within the ribosome. | '''tRNA''' or '''transfer RNA''' is stable, structured RNA present in all living cells. tRNA participates in the process of protein [[translation]] by the [[ribosome]]. Varying tRNA molecules carry a specific amino acid esterified on their 3'-OH group (the acceptor end). They also carry a specific triplet sequence, the '''anticodon''', which pairs with its complementary '''codon''' on the messenger RNA, within the ribosome. | ||
| Line 7: | Line 8: | ||
==Modified nucleotides== | ==Modified nucleotides== | ||
| - | + | ||
Most tRNAs contain modified nucleotides<ref>PMID:20459084</ref>, which are added post-transcriptionally by specific enzymes. Common modifications include isomerisation of uridines into pseudouridines (Ψ), methylation of either the ribose and/or the base, thiolation, reduction of uridines into dihydrouridines (D). The anticodon loop of the tRNA quite often contains hypermodified bases, the function of which is to stabilise the codon-anticodon interaction within the ribosome. The nature and position of nucleotide modifications is both specific of the organism and the tRNA type. | Most tRNAs contain modified nucleotides<ref>PMID:20459084</ref>, which are added post-transcriptionally by specific enzymes. Common modifications include isomerisation of uridines into pseudouridines (Ψ), methylation of either the ribose and/or the base, thiolation, reduction of uridines into dihydrouridines (D). The anticodon loop of the tRNA quite often contains hypermodified bases, the function of which is to stabilise the codon-anticodon interaction within the ribosome. The nature and position of nucleotide modifications is both specific of the organism and the tRNA type. | ||
Common modified nucleotides include : | Common modified nucleotides include : | ||
| Line 24: | Line 25: | ||
==Aminoacylation and function as an aminoacid carrier== | ==Aminoacylation and function as an aminoacid carrier== | ||
| - | < | + | <scene name='43/433638/Cv/2'>Glutaminyl-tRNA synthetase/tRNA complex</scene> ([[1gtr]]). |
Within the cell, each tRNA undergoes an aminoacylation-deacylation cycle. First, the cognate aminoacid is esterified on its 3'-OH by the cognate aminoacyl-tRNA synthetase. The synthetase recognizes structural features on the tRNA, which allows it to discriminate tRNA that are specific for a given aminoacid, from all other (non-cognate) tRNA. These structural features are called identity determinants. They are often (but not exclusively) located in the anticodon sequence and/or in the so-called discriminator base (position 73), just before the 3' -CCA terminus. | Within the cell, each tRNA undergoes an aminoacylation-deacylation cycle. First, the cognate aminoacid is esterified on its 3'-OH by the cognate aminoacyl-tRNA synthetase. The synthetase recognizes structural features on the tRNA, which allows it to discriminate tRNA that are specific for a given aminoacid, from all other (non-cognate) tRNA. These structural features are called identity determinants. They are often (but not exclusively) located in the anticodon sequence and/or in the so-called discriminator base (position 73), just before the 3' -CCA terminus. | ||
Once aminoacylated, tRNA associate with the elongation factor EF-Tu (bacteria) or EF1 (eucaryotes) complexed to GTP. These ternary complexes can then be recruited to the ribosome, where they go to the A-site. If a cognate codon-anticodon interaction is formed, translation can proceed, the aminoacid is incorporated within the polypetide chain and eventually, the deacylated tRNA is release for another aminoacylation-deacylation cycle. | Once aminoacylated, tRNA associate with the elongation factor EF-Tu (bacteria) or EF1 (eucaryotes) complexed to GTP. These ternary complexes can then be recruited to the ribosome, where they go to the A-site. If a cognate codon-anticodon interaction is formed, translation can proceed, the aminoacid is incorporated within the polypetide chain and eventually, the deacylated tRNA is release for another aminoacylation-deacylation cycle. | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | < | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==3D Structures of tRNA== | ==3D Structures of tRNA== | ||
===Free tRNA=== | ===Free tRNA=== | ||
| Line 80: | Line 74: | ||
* [[Ribosome]] | * [[Ribosome]] | ||
* [[2czj|tmRNA]] | * [[2czj|tmRNA]] | ||
| + | ==References== | ||
| + | <references/> | ||
| + | |||
| + | ==Reference for the structure== | ||
| + | <ref group="xtra">PMID:10943889</ref> | ||
| + | <references group="xtra"/> | ||
| + | |||
| + | |||
[[Category: Trna]] | [[Category: Trna]] | ||
[[Category: Topic Page]] | [[Category: Topic Page]] | ||
Revision as of 07:22, 30 July 2013
| |||||||||||
Contents |
3D Structures of tRNA
Free tRNA
yeast phenylalanine tRNA
human lysine tRNA (primer of HIV1 reverse transcription)
yeast aspartic acid tRNA
E. coli initiatior methionine tRNA
tRNA fragments
Complexes with aminoacyl-tRNA synthetases
Complexes with elongation factors
Complex with RNAse P
Complexes with the ribosome
See Also
References
- ↑ Motorin Y, Helm M. tRNA stabilization by modified nucleotides. Biochemistry. 2010 Jun 22;49(24):4934-44. PMID:20459084 doi:10.1021/bi100408z
Reference for the structure
- Shi H, Moore PB. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: a classic structure revisited. RNA. 2000 Aug;6(8):1091-105. PMID:10943889
Proteopedia Page Contributors and Editors (what is this?)
Karsten Theis, Wayne Decatur, Michal Harel, Frédéric Dardel, Ann Taylor, Joel L. Sussman, Alexander Berchansky
Categories: Trna | Topic Page | Translation | Modification | RNA | Amino acid