RNase A
From Proteopedia
| Line 1: | Line 1: | ||
| + | <StructureSection | PDB=7RSA | Size =500 | Side=right | scene ='Sandbox_Reserved_193/Rnasei_a/1' | caption='Bovine Pancreatic Ribonuclease A (RNase A), [[7rsa]]'>__NoTOC__ | ||
== '''Introduction''' == | == '''Introduction''' == | ||
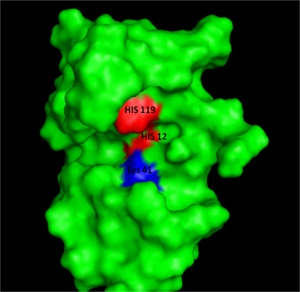
[[Image:RNaseAIII.png|300px|left|thumb|Figure I: Bovine Ribonuclease A. Colored residues are representative of amino acids important to both the acid base catalysis (Red: His12 and 119) and stabilization of the transition state (Blue: Lys41). Figure generated via ''Pymol'' ]] | [[Image:RNaseAIII.png|300px|left|thumb|Figure I: Bovine Ribonuclease A. Colored residues are representative of amino acids important to both the acid base catalysis (Red: His12 and 119) and stabilization of the transition state (Blue: Lys41). Figure generated via ''Pymol'' ]] | ||
| Line 9: | Line 10: | ||
=='''Structure, Catalysis, and Substrate Binding'''== | =='''Structure, Catalysis, and Substrate Binding'''== | ||
| - | + | ||
=='''Structure'''== | =='''Structure'''== | ||
RNase A is made up of a single polypeptide chain of 124 residues. Of the 20 natural amino acids, RNase A possesses 19 of them, excluding tryptophan.<ref name="Raines" /> This single polypeptide chain is cross-linked internally by four disulfide linkages, which contribute to the stability of RNase A. Long four-stranded anti-parallel <scene name='Sandbox_Reserved_192/Beta_sheet/4'>ß-sheets</scene> and three short <scene name='Sandbox_Reserved_192/Alpha_helices/2'>α-helices</scene> make up the <scene name='Sandbox_Reserved_192/Secondary_structure/3'>secondary structure</scene> of RNase A.<ref name="Raines" /> The amino acid sequence was discovered to determine the three-dimensional structure of RNase A by Christian Anfinsen in the 1950s. Urea was used to denature RNase A, and mercaptoethanol was used to reduce and cleave the four disulfide bonds in RNase A to yield eight Cys residues. Catalytic activity was lost due to denaturation. When the urea and mercaptoethanol were removed, the denatured ribonuclease refolded spontaneously into its correct tertiary structure with restoration of its catalytic activity. Disulfide bonds were also reformed in the same position. The Anfinsen experiment provided evidence that the amino acid sequence contained all the information required for the protein to fold into its native three-dimensional structure. Anfinsen received the 1972 Nobel Prize in Chemistry for his work with RNase A. Nevertheless, ensuing work showed some proteins require further assistance, such as molecular chaperones, to fold into their native structure.<ref name="Lehninger" /> | RNase A is made up of a single polypeptide chain of 124 residues. Of the 20 natural amino acids, RNase A possesses 19 of them, excluding tryptophan.<ref name="Raines" /> This single polypeptide chain is cross-linked internally by four disulfide linkages, which contribute to the stability of RNase A. Long four-stranded anti-parallel <scene name='Sandbox_Reserved_192/Beta_sheet/4'>ß-sheets</scene> and three short <scene name='Sandbox_Reserved_192/Alpha_helices/2'>α-helices</scene> make up the <scene name='Sandbox_Reserved_192/Secondary_structure/3'>secondary structure</scene> of RNase A.<ref name="Raines" /> The amino acid sequence was discovered to determine the three-dimensional structure of RNase A by Christian Anfinsen in the 1950s. Urea was used to denature RNase A, and mercaptoethanol was used to reduce and cleave the four disulfide bonds in RNase A to yield eight Cys residues. Catalytic activity was lost due to denaturation. When the urea and mercaptoethanol were removed, the denatured ribonuclease refolded spontaneously into its correct tertiary structure with restoration of its catalytic activity. Disulfide bonds were also reformed in the same position. The Anfinsen experiment provided evidence that the amino acid sequence contained all the information required for the protein to fold into its native three-dimensional structure. Anfinsen received the 1972 Nobel Prize in Chemistry for his work with RNase A. Nevertheless, ensuing work showed some proteins require further assistance, such as molecular chaperones, to fold into their native structure.<ref name="Lehninger" /> | ||
| Line 54: | Line 55: | ||
</StructureSection> | </StructureSection> | ||
=='''Inhibitors'''== | =='''Inhibitors'''== | ||
| - | + | <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_rnasea_simple/1'>inhibitor (RI) (tan) bound to RNase A (red)</scene> ([[1dfj]]). | |
| - | < | + | |
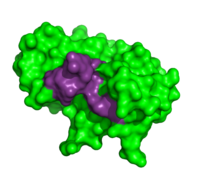
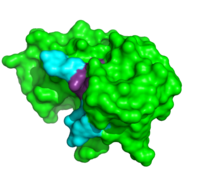
Due to the high rate of RNA hydrolysis by RNase A, mammalian cells have developed a protective inhibitor to prevent pancreatic ribonucleases from degrading cystolic RNA. Ribonuclease Inhibitor (RI) tightly associates to the active site of RNase A due to its <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_simple/1'>non-globular nature</scene>. RI is a 50 kD protein that is composed of 16 repeating subunits of alpha helices and beta sheets, giving it a noticeable <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_nonglobular/1'>horseshoe like appearance</scene>. The RI-RNase protein-protein interaction has the highest known affinity of any protein-protein interactions with an approximate dissociation constant (''K''d) of 5.8 X 10-14 for almost all types of ribonucleases.<ref>PMID:7877692</ref> The ability to be selective for almost all types of RNases, and yet retain such a high Kd is product of its mechanism of inhibition. The interior residues of the horseshoe shaped RI are able to bind to the charged residues of the active site cleft of RNase A, such as <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_rnasea_lys7_gln11_lys41/1'>Lys7, Gln11, and Lys41 </scene>. By studying the amphibian RNase, Onconase, the residues Lys7 and Gln11 of RNase A were shown to be the most important in this interaction. In onconase, these residues are replaced with non-charged amino acids, which help prevent the binding of RI to the protein <ref>PMID:18930025</ref> | Due to the high rate of RNA hydrolysis by RNase A, mammalian cells have developed a protective inhibitor to prevent pancreatic ribonucleases from degrading cystolic RNA. Ribonuclease Inhibitor (RI) tightly associates to the active site of RNase A due to its <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_simple/1'>non-globular nature</scene>. RI is a 50 kD protein that is composed of 16 repeating subunits of alpha helices and beta sheets, giving it a noticeable <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_nonglobular/1'>horseshoe like appearance</scene>. The RI-RNase protein-protein interaction has the highest known affinity of any protein-protein interactions with an approximate dissociation constant (''K''d) of 5.8 X 10-14 for almost all types of ribonucleases.<ref>PMID:7877692</ref> The ability to be selective for almost all types of RNases, and yet retain such a high Kd is product of its mechanism of inhibition. The interior residues of the horseshoe shaped RI are able to bind to the charged residues of the active site cleft of RNase A, such as <scene name='User:R._Jeremy_Johnson/RNaseA/Ri_rnasea_lys7_gln11_lys41/1'>Lys7, Gln11, and Lys41 </scene>. By studying the amphibian RNase, Onconase, the residues Lys7 and Gln11 of RNase A were shown to be the most important in this interaction. In onconase, these residues are replaced with non-charged amino acids, which help prevent the binding of RI to the protein <ref>PMID:18930025</ref> | ||
| - | + | </StructureSection> | |
| - | + | __NOTOC__ | |
| - | + | ||
| - | + | ||
=='''Additional Proteopedia Pages about RNase A'''== | =='''Additional Proteopedia Pages about RNase A'''== | ||
Revision as of 10:57, 30 July 2013
| |||||||||||
Inhibitors
(1dfj).
Due to the high rate of RNA hydrolysis by RNase A, mammalian cells have developed a protective inhibitor to prevent pancreatic ribonucleases from degrading cystolic RNA. Ribonuclease Inhibitor (RI) tightly associates to the active site of RNase A due to its . RI is a 50 kD protein that is composed of 16 repeating subunits of alpha helices and beta sheets, giving it a noticeable . The RI-RNase protein-protein interaction has the highest known affinity of any protein-protein interactions with an approximate dissociation constant (Kd) of 5.8 X 10-14 for almost all types of ribonucleases.[11] The ability to be selective for almost all types of RNases, and yet retain such a high Kd is product of its mechanism of inhibition. The interior residues of the horseshoe shaped RI are able to bind to the charged residues of the active site cleft of RNase A, such as . By studying the amphibian RNase, Onconase, the residues Lys7 and Gln11 of RNase A were shown to be the most important in this interaction. In onconase, these residues are replaced with non-charged amino acids, which help prevent the binding of RI to the protein [12] </StructureSection>
Additional Proteopedia Pages about RNase A
3D structures of ribonuclease
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 1.9 Raines RT. Ribonuclease A. Chem Rev. 1998 May 7;98(3):1045-1066. PMID:11848924
- ↑ Avey HP, Boles MO, Carlisle CH, Evans SA, Morris SJ, Palmer RA, Woolhouse BA, Shall S. Structure of ribonuclease. Nature. 1967 Feb 11;213(5076):557-62. PMID:6032249
- ↑ Wyckoff HW, Hardman KD, Allewell NM, Inagami T, Johnson LN, Richards FM. The structure of ribonuclease-S at 3.5 A resolution. J Biol Chem. 1967 Sep 10;242(17):3984-8. PMID:6037556
- ↑ Greenway MJ, Andersen PM, Russ C, Ennis S, Cashman S, Donaghy C, Patterson V, Swingler R, Kieran D, Prehn J, Morrison KE, Green A, Acharya KR, Brown RH Jr, Hardiman O. ANG mutations segregate with familial and 'sporadic' amyotrophic lateral sclerosis. Nat Genet. 2006 Apr;38(4):411-3. Epub 2006 Feb 26. PMID:16501576 doi:10.1038/ng1742
- ↑ 5.0 5.1 5.2 'Lehninger A., Nelson D.N, & Cox M.M. (2008) Lehninger Principles of Biochemistry. W. H. Freeman, fifth edition.'
- ↑ 6.0 6.1 6.2 Wlodawer A, Svensson LA, Sjolin L, Gilliland GL. Structure of phosphate-free ribonuclease A refined at 1.26 A. Biochemistry. 1988 Apr 19;27(8):2705-17. PMID:3401445
- ↑ Birdsall DL, McPherson A. Crystal structure disposition of thymidylic acid tetramer in complex with ribonuclease A. J Biol Chem. 1992 Nov 5;267(31):22230-6. PMID:1429575
- ↑ delCardayre SB, Raines RT. Structural determinants of enzymatic processivity. Biochemistry. 1994 May 24;33(20):6031-7. PMID:8193116
- ↑ Thompson JE, Raines RT. Value of general Acid-base catalysis to ribonuclease a. J Am Chem Soc. 1994 Jun;116(12):5467-8. PMID:21391696 doi:10.1021/ja00091a060
- ↑ Fontecilla-Camps JC, de Llorens R, le Du MH, Cuchillo CM. Crystal structure of ribonuclease A.d(ApTpApApG) complex. Direct evidence for extended substrate recognition. J Biol Chem. 1994 Aug 26;269(34):21526-31. PMID:8063789
- ↑ Kobe B, Deisenhofer J. A structural basis of the interactions between leucine-rich repeats and protein ligands. Nature. 1995 Mar 9;374(6518):183-6. PMID:7877692 doi:http://dx.doi.org/10.1038/374183a0
- ↑ Turcotte RF, Raines RT. Interaction of onconase with the human ribonuclease inhibitor protein. Biochem Biophys Res Commun. 2008 Dec 12;377(2):512-4. Epub 2008 Oct 16. PMID:18930025 doi:10.1016/j.bbrc.2008.10.032
External Resources
- Full crystal structure of 1RTA
- Full crystal structure of 1RCN
- Further information on 1RTA
- RNase A- Molecule of the Month
- RNase A-Wikipedia
- Ribonuclease Inhibitor-Wikipedia
- Ribonuclease-Wikipedia
- Ruminants-Wikipedia
Student Contributors
- Nathan Clarke and Kristyn Shaw
- Mary Andorfer, Grace Douglass, and Laurel Heckman
- Lauren Garnett and Micah Raebel
Proteopedia Page Contributors and Editors (what is this?)
Alexander Berchansky, R. Jeremy Johnson, Karsten Theis, Angel Herraez, Michal Harel