Introduction
(TP), also known as penicillin-binding proteins
(PBP), catalyze the cross-linking of peptidoglycan polymers during
bacterial cell wall synthesis. Beta-lactam (β-lactam) antibiotics,
which include penicillins, cephalosporins and carbapenems, bind and
irreversibly inhibit transpeptidases. The overuse and misuse of β-lactam
antibiotics has led to strains of Staphylococcus aureus that are resistant
to all β-lactams and are often only susceptible to “last resort antibiotics”,
such as vancomycin.
Cell Wall Structure
The cell wall, which is composed of peptidoglycans, is crucial for maintaining
the structural integrity of the bacterium. Peptidoglycans consists of
N-acetylmuramic Acid (NAM) and N-acetylglucosamine (NAG) polymers. Rows of
peptidoglycan are cross-linked together with pentaglycine chains. The NAM residues
have a five amino acid side chain that terminates with two D-Alanine (D-Ala)
residues.
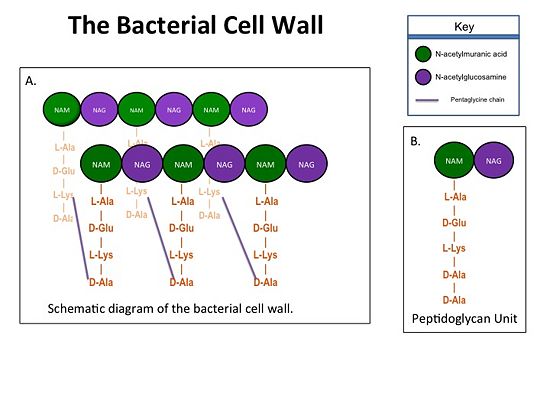
Figure 1.(A)Rows of Peptidoglycans forming a Bacterial Cell Wall (B)Peptidoglycan with D-Ala-D-Ala substrate
Structure of a Resistant Transpeptidase
Methicillin resistant Staphylococcus aureus (MRSA) is resistant to all β-lactams because it acquires an alternative PBP, PBP2a, that is not bound or inhibited by any β-lactams. PBP2a is composed of two domains:
(NPB) domain and a domain. The NBP domain of PBP2a is anchored in the cell membrane, while the TP domain “sits” in the periplasm with its active site facing the inner surface of the cell wall. The active site contains a serine residue at position 403 ()which catalyzes the cross-linking of the peptidoglycan rows with pentaglycine cross-links.
Catalytic Mechanism of PBP2a
(a) The D-Ala-D-Ala side-chain substrate of the peptidoglycan accesses
the active site of the PBP2a.
(b) Ser403 nucleophilically attacks the peptide bond of the terminal
D-Ala residues of the substrate. The terminal D-Ala residue then exits
the active site. The now terminal D-Ala residue forms a covalent bond to Ser403,
while a crosslinking pentaglycine chain enters the active site.
(c) A covalent bond forms between the pentaglycine chain and the
terminal D-Ala residue, regenerating the active site serine residue.
The entire process takes 4 milliseconds.
How do antibiotics work?
The β-lactam antibiotics inhibit bacterial growth by inhibiting PBPs and ultimately cell wall
synthesis. Specifically, β-lactams are molecular mimics of D-Ala-D-Ala, which is the normal
substrate of PBPs. Nucleophillic attack of the β-lactam results in the PBP being irreversibly
inhibited by the β-lactam. As a result, the synthesis of the cell wall is inhibited which leads
to cell lysis.
PBP2a and Ceftobiprole
MRSA becomes resistant to β-lactams by acquiring an alternative PBP, PBP2a, that is
neither bound nor inhibited by β-lactams. Recently, two cephalosporins –
and
ceftaroline – that have anti-MRSA activity have been developed. Ceftobiprole is able to
inhibit PBP2a because additional chemical groups at the
position of the cephalosporin backbone are able to interact with additional amino acid
residues in PBP2a; specifically .
As a result of its tighter binding to PBP2a,
is able to more efficiently react with the serine active site residue and therefore inhibit the activity of PBP2a.

