This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox 127
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
<StructureSection load= size=475 side='right' scene='36/365380/4dki_cartoon/22'> | <StructureSection load= size=475 side='right' scene='36/365380/4dki_cartoon/22'> | ||
| - | Peptidoglycan | + | Peptidoglycan transpeptidases (TP), also known as penicillin-binding proteins (PBP), are essential for bacterial cell wall synthesis and catalyze the cross-linking of peptidoglycan polymers during bacterial cell wall synthesis. Beta-lactam (β-lactam) antibiotics, which include the penicillins, cephalosporins, carbapenems, and the monobactam aztreonam (Figure 1); bind and irreversibly inhibit the active site of TP. The overuse and misuse of β-lactam antibiotics has led to strains of Staphylococcus aureus (S. aureus) that are resistant to almost all currently available β-lactams and are often only susceptible to so-called “last resort antibiotics”, such as vancomycin. |
| + | |||
| + | |||
== Cell Wall Structure == | == Cell Wall Structure == | ||
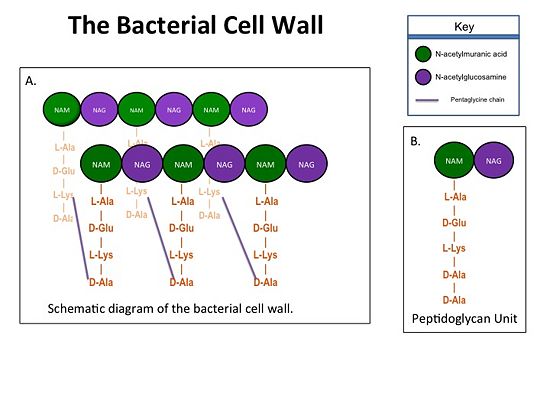
| - | + | The bacterial cell wall is crucial for maintaining the structural integrity of bacteria and protects bacteria from osmotic stress and toxic compounds. The cell wall is composed of peptidoglycan (Figure 2 ) and in Gram positive bacterial species (e.g., S. aureus) is many layers thick, while in Gram negative bacterial species (e.g., Escherichia coli) is only a few layers thick. The difference in the number of peptidoglycan layers accounts for the differential staining of these two groups of organisms. . Peptidoglycan consists of a carbohydrate portion: alternating residues of N-acetylmuranic Acid (NAM) and N-acetylglucosamine (NAG) that polymerize to form long chains, and a protein portion: a pentapeptide chain that terminates with two D-alanine (D-Ala) residues. The pentapeptide chains are covalently bound to each NAM residue Rows of peptidoglycan are cross-linked together with pentaglycine chains to form a "mesh-like" structure. This cross-linking reaction is catalyzed by TPs. | |
| - | The cell wall | + | |
[[Image:Cell Wall 7 30 2013.jpg|thumb|alt= Alt text| Figure 1. A.Bacterial Cell Wall B.Peptidoglycan with D-Ala-D-Ala substrate |550px]] | [[Image:Cell Wall 7 30 2013.jpg|thumb|alt= Alt text| Figure 1. A.Bacterial Cell Wall B.Peptidoglycan with D-Ala-D-Ala substrate |550px]] | ||
| Line 15: | Line 16: | ||
| - | == Catalytic Mechanism of | + | == Catalytic Mechanism of Action of Transpeptidases == |
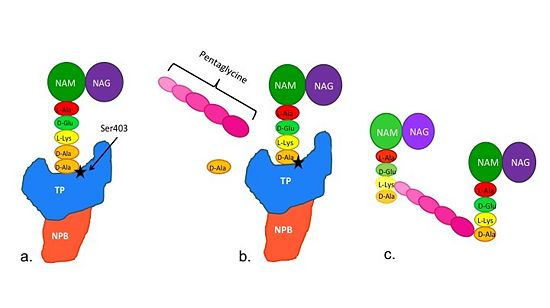
[[Image:Schematic TP 3steps.jpg|thumb|alt= Alt text|Figure 2. Schematic diagram illustrating the mechanism of action of PBP2a |550px]] | [[Image:Schematic TP 3steps.jpg|thumb|alt= Alt text|Figure 2. Schematic diagram illustrating the mechanism of action of PBP2a |550px]] | ||
| + | Note: Schematic is above these statements: | ||
| + | (a)The D-Ala-D-Ala side-chain substrate accesses the TP active site. | ||
| + | (b) The active site serine residue nucleophilically attacks the peptide bond between the terminal D-Ala residues. The terminal D-Ala residue exits the active site, and the remaining D-Ala residue forms a covalent bond with the active site serine residue to form an acyl-TP complex. | ||
| + | (c) Subsequently, a pentaglycine chain enters the TP active site through nucleophillic attack forms a covalent bond with the D-Ala residue formerly bound to the active site serine residue. As a result, the TP active site serine residue is regenerated. The entire process takes approximately 4 milliseconds. | ||
| - | (a)The D-Ala-D-Ala side-chain substrate of the peptidoglycan accesses the active site of the PBP2a. | ||
| - | |||
| - | (b)Ser403 nucleophilically attacks the peptide bond of the terminal D-Ala residues of the substrate. The terminal D-Ala residue then exits the active site. The now terminal D-Ala residue forms a covalent bond to Ser403, while a crosslinking pentaglycine chain enters the active site. | ||
| - | |||
| - | (c)A covalent bond forms between the pentaglycine chain and the terminal D-Ala residue, regenerating the active site serine residue. | ||
| - | + | == Mechanism of Action of β-Lactam Antibiotics == | |
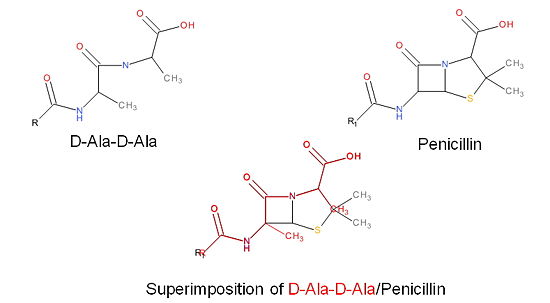
| - | + | The β-lactam antibiotics inhibit bacterial growth by irreversibly inhibiting TPs and, therefore, bacterial cell wall synthesis. Specifically, β-lactams are molecular mimics of a portion of the peptidoglycan polymer, namely the D-Ala-D-Ala moiety, which is the normal TP enzymatic substrate (Figure 4). As a result, bacterial TP enzymes are "tricked" into reacting with β-lactams. Additionally, the β-lactams are very reactive molecules due to their β-lactam ring, and readily react with the TP active site serine residue and sterically block the active site preventing the entry of nucleophiles [http://en.wikipedia.org/wiki/Nucleophile] that regenerate the active site serine residue such as the pentaglycine chain or water. | |
| - | The β-lactam antibiotics inhibit bacterial growth by inhibiting PBPs and ultimately cell wall | ||
| - | synthesis. Specifically, β-lactams are molecular mimics of D-Ala-D-Ala, which is the normal | ||
| - | substrate of PBPs. Nucleophillic attack [http://en.wikipedia.org/wiki/Nucleophile] of the β-lactam results in the PBP being irreversibly inhibited by the β-lactam. As a result, the synthesis of the cell wall is inhibited which leads to cell lysis. | ||
[[Image:Structures on penicillin and b lactam.jpg|thumb|alt= Alt text|Figure 3. Mechanism of action of β-lactams. A. Structure of a β-lactam (penicillin) showing the amide, carboxyl, and β-lactam ring groups β-lactam ring groups. B. Structure of the D-Ala-D-Ala substrate. C. Overlay of the D-Ala-D-Ala substrate in red with penicillin demonstrating molecular mimicry.|550 px]] | [[Image:Structures on penicillin and b lactam.jpg|thumb|alt= Alt text|Figure 3. Mechanism of action of β-lactams. A. Structure of a β-lactam (penicillin) showing the amide, carboxyl, and β-lactam ring groups β-lactam ring groups. B. Structure of the D-Ala-D-Ala substrate. C. Overlay of the D-Ala-D-Ala substrate in red with penicillin demonstrating molecular mimicry.|550 px]] | ||
Revision as of 21:18, 11 September 2013
| |||||||||||