User:Andrew Wills/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
===CRYSTAL STRUCTURE OF E. COLI 3-METHYLADENINE DNA GLYCOSYLASE (ALKA) COMPLEXED WITH DNA=== | ===CRYSTAL STRUCTURE OF E. COLI 3-METHYLADENINE DNA GLYCOSYLASE (ALKA) COMPLEXED WITH DNA=== | ||
| - | + | ||
| - | ==General Function== | + | ==Your Heading Here (maybe something like 'Structure')==<StructureSection load='1dq8' size='500' side='right' caption='Structure of HMG-CoA reductase (PDB entry [[1dq8]])' scene=''>==General Function== |
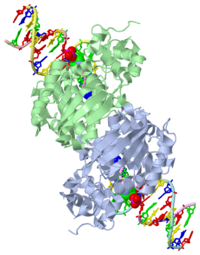
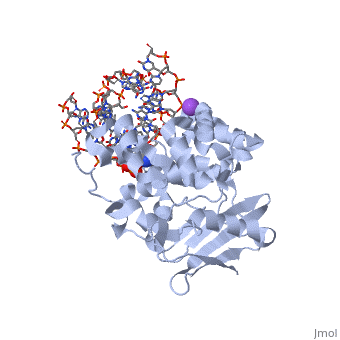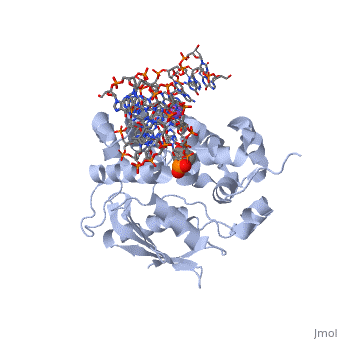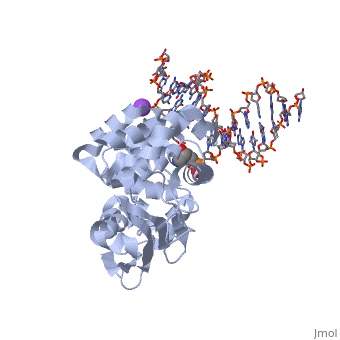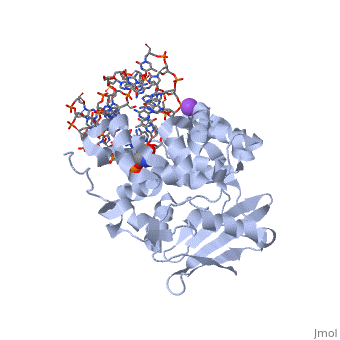
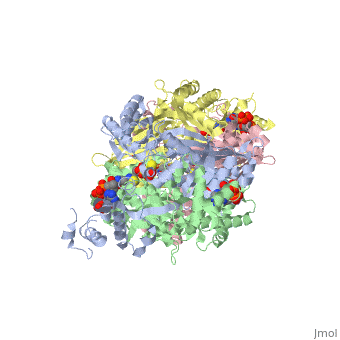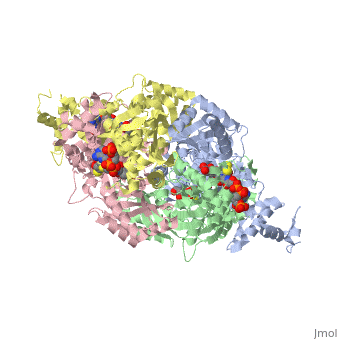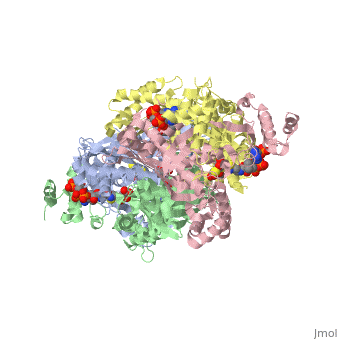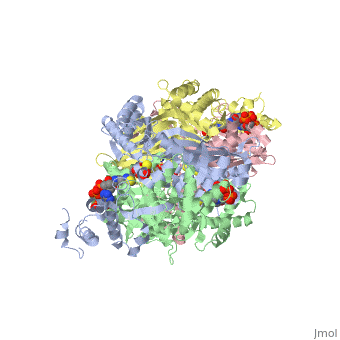
[[1diz]] Escherichia coli 3 methyladenine DNA glycosylase II (AlkA) is a DNA repair enzyme that initiates base excision repair for the removal of alkylated bases. Aflatoxin B1 is an example of a toxin that attacks guanine and adenine at their N-7 atom to form alkylated bases (stryker), which prevent regulatory proteins from binding to DNA and blocks replicative polymerases (Hollis). AlkA initiates base excision repair by first locating and binding to the alkylated DNA. It then flips the affected base out of the DNA double helix and into the active site of the enzyme. Once in the active site, AlkA hydrolyzes the glycosidic bond to release the damaged base and leave the sugar phosphate backbone intact. This creates the AP site that is either devoid of a purine or pyridine. The AP site signals to other base excision repair enzymes to insert an undamaged nucleotide based on the undamaged complementary strand and seal the DNA. | [[1diz]] Escherichia coli 3 methyladenine DNA glycosylase II (AlkA) is a DNA repair enzyme that initiates base excision repair for the removal of alkylated bases. Aflatoxin B1 is an example of a toxin that attacks guanine and adenine at their N-7 atom to form alkylated bases (stryker), which prevent regulatory proteins from binding to DNA and blocks replicative polymerases (Hollis). AlkA initiates base excision repair by first locating and binding to the alkylated DNA. It then flips the affected base out of the DNA double helix and into the active site of the enzyme. Once in the active site, AlkA hydrolyzes the glycosidic bond to release the damaged base and leave the sugar phosphate backbone intact. This creates the AP site that is either devoid of a purine or pyridine. The AP site signals to other base excision repair enzymes to insert an undamaged nucleotide based on the undamaged complementary strand and seal the DNA. | ||
| Line 21: | Line 21: | ||
Atomic Level | Atomic Level | ||
| - | In order to keep the DNA strand bound to the protein, AlkA depends on van der Waals interactions on the minor groove of DNA. The van der Waals interactions play a major role in AlkA’s preference for double stranded DNA (Hollis). An example of these interactions may be seen with the Pro175 wedged into the minor groove of DNA and anchoring it by van der Waals interctions (Hallis). | + | In order to keep the DNA strand bound to the protein, AlkA depends on van der Waals interactions on the minor groove of DNA. The van der Waals interactions play a major role in AlkA’s preference for double stranded DNA (Hollis). An example of these interactions may be seen with the Pro175 wedged into the minor groove of DNA and anchoring it by van der Waals interctions (Hallis).</StructureSection> |
| - | </StructureSection> | + | |
==See Also== | ==See Also== | ||
*[[DNA glycosylate|DNA glycosylate]] | *[[DNA glycosylate|DNA glycosylate]] | ||
Revision as of 20:52, 31 October 2013
| |||||||||
| 1diz, resolution 2.50Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Non-Standard Residues: | |||||||||
| Activity: | DNA-3-methyladenine glycosylase II, with EC number 3.2.2.21 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
CRYSTAL STRUCTURE OF E. COLI 3-METHYLADENINE DNA GLYCOSYLASE (ALKA) COMPLEXED WITH DNA
==Your Heading Here (maybe something like 'Structure')==
| |||||||||||
See Also
Reference
- Hollis T, Ichikawa Y, Ellenberger T. DNA bending and a flip-out mechanism for base excision by the helix-hairpin-helix DNA glycosylase, Escherichia coli AlkA. EMBO J. 2000 Feb 15;19(4):758-66. PMID:10675345 doi:http://dx.doi.org/10.1093/emboj/19.4.758