This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Single stranded binding protein
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | + | ===Sandbox Single Stranded DNA-Binding Protein (SSB)=== | |
| - | ==Sandbox Single Stranded DNA-Binding Protein (SSB)== | + | |
==Introduction== | ==Introduction== | ||
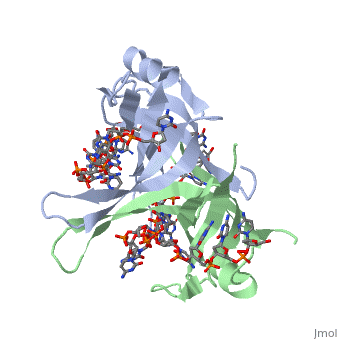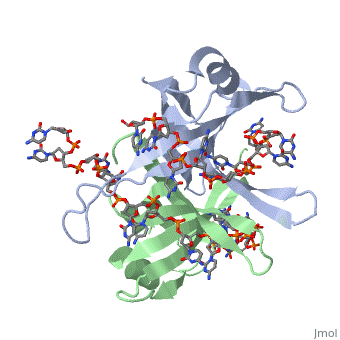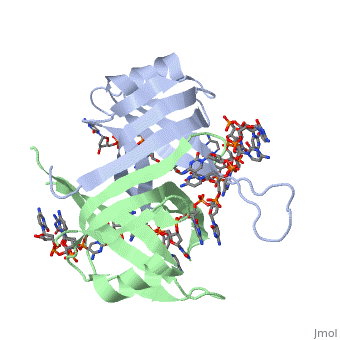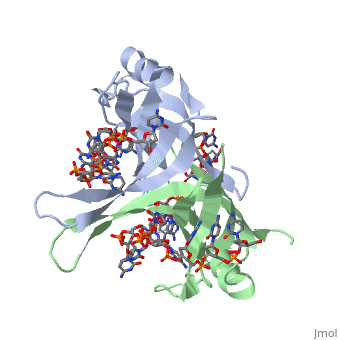
<StructureSection load='1eyg' size='400' side='right' frame='true' caption='Structure of Single Stranded DNA-Binding Protein bound to ssDNA (PDB entry [[1eyg]])' scene=''> | <StructureSection load='1eyg' size='400' side='right' frame='true' caption='Structure of Single Stranded DNA-Binding Protein bound to ssDNA (PDB entry [[1eyg]])' scene=''> | ||
| Line 12: | Line 11: | ||
</StructureSection> | </StructureSection> | ||
| - | + | ==Structure== | |
<StructureSection load='2vw9' size='400' side='right' frame='true' caption='Structure of Single Stranded DNA-Binding Protein from Helicobacter Pylori bound to ssDNA (PDB entry [[2vw9]])' scene=''> | <StructureSection load='2vw9' size='400' side='right' frame='true' caption='Structure of Single Stranded DNA-Binding Protein from Helicobacter Pylori bound to ssDNA (PDB entry [[2vw9]])' scene=''> | ||
| - | |||
| - | ==Structure== | ||
SSB consists is a homotetramer that has a DNA binding domain which binds to a single strand of DNA. The tetramers consist of α-helices, β-sheets, and random coils. Each subunit contains an <scene name='56/566528/Alpha_helices/3'>α-helix</scene> and several <scene name='56/566528/Beta_sheets/1'>β-sheets</scene>. The secondary structure also includes a NH2 terminus, which consists of multiple positively charged amino acids. The COOH terminus includes many acidic amino acids. | SSB consists is a homotetramer that has a DNA binding domain which binds to a single strand of DNA. The tetramers consist of α-helices, β-sheets, and random coils. Each subunit contains an <scene name='56/566528/Alpha_helices/3'>α-helix</scene> and several <scene name='56/566528/Beta_sheets/1'>β-sheets</scene>. The secondary structure also includes a NH2 terminus, which consists of multiple positively charged amino acids. The COOH terminus includes many acidic amino acids. | ||
Revision as of 02:18, 2 November 2013
Contents |
Sandbox Single Stranded DNA-Binding Protein (SSB)
Introduction
| |||||||||||
Structure
| |||||||||||
| |||||||||||
See Also
References
- ↑ Meyer RR, Laine PS. The single-stranded DNA-binding protein of Escherichia coli. Microbiol Rev. 1990 Dec;54(4):342-80. PMID:2087220
- ↑ Meyer RR, Laine PS. The single-stranded DNA-binding protein of Escherichia coli. Microbiol Rev. 1990 Dec;54(4):342-80. PMID:2087220
- ↑ Meyer RR, Laine PS. The single-stranded DNA-binding protein of Escherichia coli. Microbiol Rev. 1990 Dec;54(4):342-80. PMID:2087220
Proteopedia Page Contributors and Editors (what is this?)
Refayat Ahsen, Rachel Craig, Michal Harel, Alexander Berchansky