This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Andrew Wills/Sandbox 1
From Proteopedia
| Line 12: | Line 12: | ||
==Key Structures== | ==Key Structures== | ||
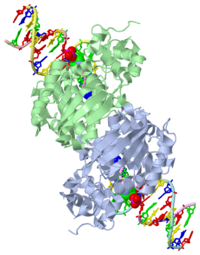
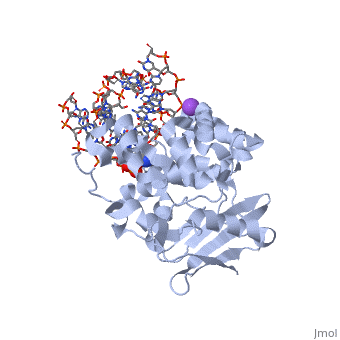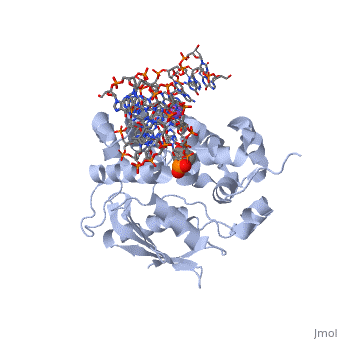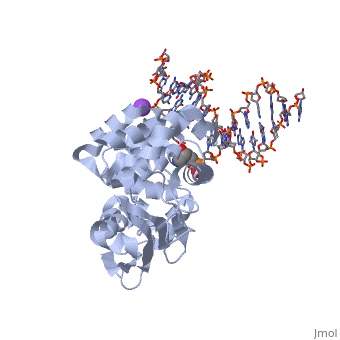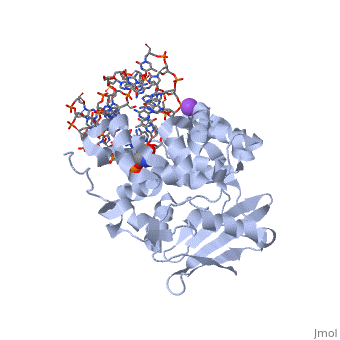
| - | AlkA is composed of three main domains with dimensions of approximately 50 Angstroms, 45 Angstroms, and 25 Angstroms.<ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> The first domain (residues 1-112) is the <scene name='56/566536/N_terminal_domain_and_dna/1'>N-terminal domain</scene> that is composed of a five stranded antiparallel beta sheet and two alpha helices.<ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> The <scene name='56/566536/Second_domain/ | + | AlkA is composed of three main domains with dimensions of approximately 50 Angstroms, 45 Angstroms, and 25 Angstroms.<ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> The first domain (residues 1-112) is the <scene name='56/566536/N_terminal_domain_and_dna/1'>N-terminal domain</scene> that is composed of a five stranded antiparallel beta sheet and two alpha helices.<ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> The <scene name='56/566536/Second_domain/2'>second domain</scene> (residues 113-230) contains seven alpha helices that create a hydrophobic core.<ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> The third domain (resudues 231-282) is the <scene name='56/566536/C-terminal/1'>C-Terminal</scene> domain that contains a bundle of three alpha helices. <ref name="Labahn">Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.</ref> |
AlkA is a member of the helix-hairpin-helix (HhH) family of DNA glycosylases, where two compact alpha helical structures are connected by a hairpin loop.<ref name="Moe">Moe, E, D.R. Hall, et al. "Structure-function studies of an unusual 3-methyladenine DNA glycosylase II (AlkA) from Deinococcus radiodurans." Biological Crystallography. 68.6 (2012): 703-712. Print. </ref> In AlkA, the <scene name='56/566536/Hhh_domain/2'>HhH</scene> domain is composed of residues 202-227 and is responsible for binding the damaged DNA by van der Waals interactions, a few hydrogen bonds, and metal ion interactions. | AlkA is a member of the helix-hairpin-helix (HhH) family of DNA glycosylases, where two compact alpha helical structures are connected by a hairpin loop.<ref name="Moe">Moe, E, D.R. Hall, et al. "Structure-function studies of an unusual 3-methyladenine DNA glycosylase II (AlkA) from Deinococcus radiodurans." Biological Crystallography. 68.6 (2012): 703-712. Print. </ref> In AlkA, the <scene name='56/566536/Hhh_domain/2'>HhH</scene> domain is composed of residues 202-227 and is responsible for binding the damaged DNA by van der Waals interactions, a few hydrogen bonds, and metal ion interactions. | ||
Revision as of 14:57, 6 November 2013
| |||||||||
| 1diz, resolution 2.50Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Non-Standard Residues: | |||||||||
| Activity: | DNA-3-methyladenine glycosylase II, with EC number 3.2.2.21 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
CRYSTAL STRUCTURE OF E. COLI 3-METHYLADENINE DNA GLYCOSYLASE (ALKA) COMPLEXED WITH DNA
<nowiki>Insert non-formatted text here</nowiki>
General Function
1diz Escherichia coli 3 methyladenine DNA glycosylase II () is a DNA repair enzyme that initiates base excision repair for the removal of alkylated bases. Aflatoxin B1 is an example of a toxin that attacks guanosine at its N-7 atom to form alkylated bases[1], which prevent regulatory proteins from binding to DNA and blocks replicative polymerases.[2] AlkA initiates base excision repair by first locating and binding to the alkylated DNA. It then flips the affected base out of the DNA double helix and into the active site of the enzyme. Once in the active site, AlkA hydrolyzes the glycosidic bond to release the damaged base and leave the sugar phosphate backbone intact. This creates the AP site that is either devoid of a purine or pyridine. The AP site signals to other base excision repair enzymes to insert an undamaged nucleotide based on the undamaged complementary strand and seal the DNA.
| |||||||||||
See Also
References
- ↑ Berg, Jeremy, Tymoczko John, and Lubert Stryer. Biochemistry. 6th. New York: W.H. Freeman and Company, 2007. 806-808. Print.
- ↑ 2.0 2.1 2.2 Hollis, Thomas, Yoshitaka Ichikawa, and Tom Ellenberger. "DNA bending and a flip-out mechanism for base excision by the helix-hairpin-helix DNA glycosylase, Escherichia coli AlkA." EMBO Journal. 19.4 (2000): 758-766. Print.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Labahn, Jorg, Orlando Scharer, et al. "Structural Basis for the Excision Repair of Alkylation-Damaged DNA." Cell. 86.2 (1996): 321-329. Print.
- ↑ 4.0 4.1 4.2 4.3 Moe, E, D.R. Hall, et al. "Structure-function studies of an unusual 3-methyladenine DNA glycosylase II (AlkA) from Deinococcus radiodurans." Biological Crystallography. 68.6 (2012): 703-712. Print.