This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
ATP-dependent DNA ligase from bacteriophage T7
From Proteopedia
(Difference between revisions)
| Line 18: | Line 18: | ||
==Catalytic function with DNA== | ==Catalytic function with DNA== | ||
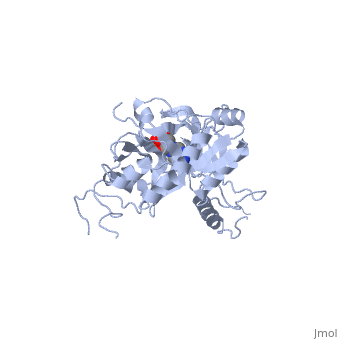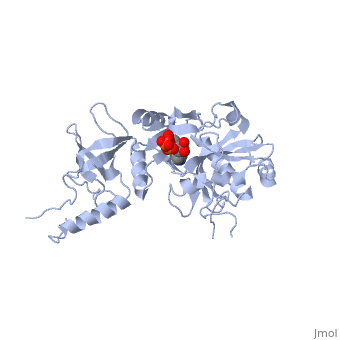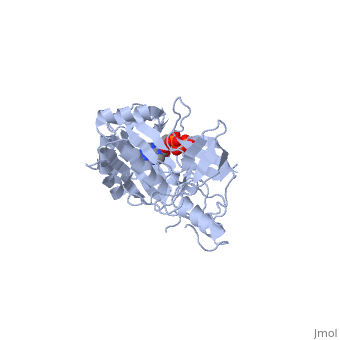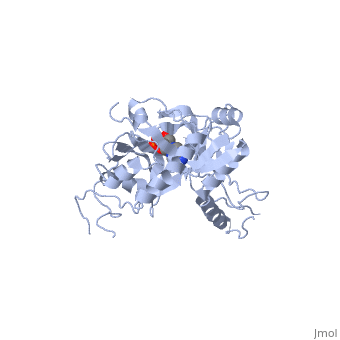
| - | ATP-dependent DNA ligase from bacteriophage T7 amends a fractured DNA strand through esterification of a 5'- phosphoryl to a 3'- hydroxyl group<ref name="Bacteriophage"/>. This mechanism occurs with the aid of ATP in several steps. First, the ligase is activated through a <scene name='56/567310/Amp_complex/1'>phosphoramidate bond with a lysine residue</scene> in the active site (Lys 34). A pyrophosphate leaves and the enzyme-AMP complex is formed. Next, the AMP is transferred to the 5' phosphate group at the nick in the DNA. Finally, T7 ligase creates the phosphodiester bond between the 5' -phosphoryl and the 3' – hydroxyl group, with AMP being freed<ref name="Crystal"/>. All ATP-dependent DNA ligases contain a conserved amino acid sequence of KxDGxR<ref name="Bacteriophage"/>. This includes the lysine residue which binds the ATP in the groove between the two domains. | + | ATP-dependent DNA ligase from bacteriophage T7 amends a fractured DNA strand through esterification of a 5'- phosphoryl to a 3'- hydroxyl group<ref name="Bacteriophage"/>. This mechanism occurs with the aid of ATP in several steps. First, the ligase is activated through a <scene name='56/567310/Amp_complex/1'>phosphoramidate bond with a lysine residue</scene> in the active site (Lys 34). A pyrophosphate leaves and the enzyme-AMP complex is formed. Next, the AMP is transferred to the 5' phosphate group at the nick in the DNA. Finally, T7 ligase creates the phosphodiester bond between the 5' -phosphoryl and the 3' – hydroxyl group, with AMP being freed<ref name="Crystal"/>. All ATP-dependent DNA ligases contain a conserved amino acid sequence of <scene name='56/567310/Residues_34-39/1'>KxDGxR</scene><ref name="Bacteriophage"/>. This includes the lysine residue which binds the ATP in the groove between the two domains. |
</StructureSection> | </StructureSection> | ||
Revision as of 07:03, 7 November 2013
ATP-DEPENDENT DNA LIGASE FROM BACTERIOPHAGE T7
| |||||||||||
References
- ↑ McGrath S, van Sinderen D. Bacteriophage: Genetics and Molecular Biology. Norfolk: Caister Academic Press, 2007. Print.
- ↑ Berg, Jeremy M, Stryer, Lubert, Tymoczko, John L. Biochemistry. Sixth edition. New York: W.H. Freeman and Company, 2007: 796. Print.
- ↑ 3.0 3.1 3.2 Doherty AJ, Ashford SR, Subramanya HS, Wigley DB. Bacteriophage T7 DNA ligase. Overexpression, purification, crystallization, and characterization. J Biol Chem. 1996 May 10;271(19):11083-9. PMID:8626651
- ↑ 4.0 4.1 4.2 4.3 4.4 Subramanya HS, Doherty AJ, Ashford SR, Wigley DB. Crystal structure of an ATP-dependent DNA ligase from bacteriophage T7. Cell. 1996 May 17;85(4):607-15. PMID:8653795
Proteopedia Page Contributors and Editors (what is this?)
William Guthrie, Hunter Douglas, Jeremy A. Hammett, Jaime Prilusky