This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Prion
From Proteopedia
| Line 1: | Line 1: | ||
| + | <StructureSection load='3haf' size='450' side='right' scene='Prion/Cv/3' caption=''> | ||
[[Image:3haf.png|left|200px|thumb|Crystal Structure of human prion [[3haf]]]] | [[Image:3haf.png|left|200px|thumb|Crystal Structure of human prion [[3haf]]]] | ||
| - | {{ | + | {{Clear}} |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
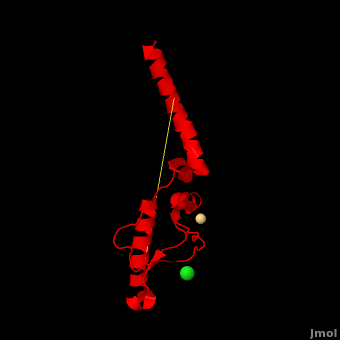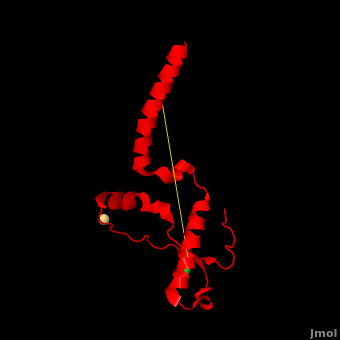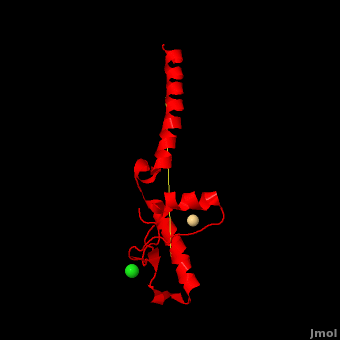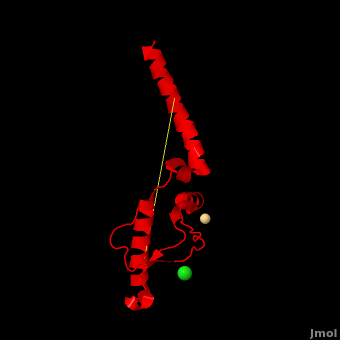
'''Prion''' (PrP) is a protein which becomes infectious upon undergoing conformation change to an amyloid form, which is self-propagating and becomes resistant to protease degradation. The fungus ''Podospora anserine'' has a prion-like protein HET-S which undergoes a conformation change to amyloid form which prevents its colony from merging with non-compatible colonies. Yeast prion proteins are Sup35 and Ure2. The images at the left and at the right correspond to one representative prion, ''i.e.'' the crystal structure of human prion ([[3haf]]). For more details see<br /> | '''Prion''' (PrP) is a protein which becomes infectious upon undergoing conformation change to an amyloid form, which is self-propagating and becomes resistant to protease degradation. The fungus ''Podospora anserine'' has a prion-like protein HET-S which undergoes a conformation change to amyloid form which prevents its colony from merging with non-compatible colonies. Yeast prion proteins are Sup35 and Ure2. The images at the left and at the right correspond to one representative prion, ''i.e.'' the crystal structure of human prion ([[3haf]]). For more details see<br /> | ||
*[[Prion protein]]<br /> | *[[Prion protein]]<br /> | ||
*[[Human Prion Protein Dimer]].<br /> | *[[Human Prion Protein Dimer]].<br /> | ||
| - | Click here to see <scene name='Prion/Cv/2'>example of prion unfolding</scene> (morph was taken from [http://molmovdb.org/cgi-bin/movie.cgi Gallery of Morphs] of the [http://molmovdb.org Yale Morph Server]). | + | Click here to see <scene name='Prion/Cv/2'>example of prion unfolding</scene> (morph was taken from [http://molmovdb.org/cgi-bin/movie.cgi Gallery of Morphs] of the [http://molmovdb.org Yale Morph Server]). |
| + | === Dominant-negative Effects in Prion Diseases: Insights from Molecular Dynamics Simulations on <scene name='Journal:JBSD:4/Cv/2'>Mouse Prion Protein Chimeras</scene> <ref>doi 10.1080/07391102.2012.712477</ref>=== | ||
| + | The key event in prion diseases is the conformational conversion from the cellular form of the [[Prion_protein|prion protein]] (PrP<sup>C</sup>) to its pathogenic scrapie form PrP<sup>Sc</sup> (or prion). PrP<sup>Sc</sup> is the sole causative agent of prion diseases which self-propagates by converting PrP<sup>C</sup> to nascent PrP<sup>Sc</sup>. Mutations in the open reading sequence of the [[Prion_protein|prion protein]] gene can introduce changes in the protein structure and alter PrP<sup>Sc</sup> formation and propagation, possibly by (de)stabilizing the physiological folding of PrP<sup>C</sup> and/or affecting its interactions with some yet unknown cellular factors. Some PrP polymorphisms may even inhibit the wild-type (WT) PrP<sup>C</sup> from being converted to PrP<sup>Sc</sup>, with the so-called “dominant-negative” effect. | ||
| + | Here we use molecular dynamics simulations to investigate the structural determinants of the globular domain in engineered Mouse (Mo) PrP variants, in WT human (Hu) PrP (PDB: [[1hjn]]) and in WT MoPrP (PDB: [[1xyx]]). The Mo PrP variants investigated here contain one or two residues from ''Homo sapiens'' and are denoted “MoPrP chimeras”. <scene name='Journal:JBSD:4/Cv/3'>Some of them are resistant to PrP<sup>Sc</sup> infection</scene> <span style="color:yellow;background-color:black;font-weight:bold;">(colored in yellow)</span> in ''in vivo'' or in ''in vitro'' cell-culture experiments, the <scene name='Journal:JBSD:4/Cv/7'>others are not</scene> <font color='darkmagenta'><b>(in darkmagenta)</b></font>. Our main results are the following: (i) The chimeras resistant to PrP<sup>Sc</sup> infection show <scene name='Journal:JBSD:4/Cv/8'>shorter intramolecular distances</scene> between the α1 helix and N-terminal of α3 helix than HuPrP, MoPrP and the non-resistant chimeras (<scene name='Journal:JBSD:4/Cv/12'>click here to see morph</scene>). This is due to stronger specific interactions between these two regions, mainly the <scene name='Journal:JBSD:4/Cv/9'>Y149-D202 and D202-Y157 (in Hu numbering and hereafter) hydrogen bonds</scene> and the <scene name='Journal:JBSD:4/Cv/10'>R156-E196 salt bridge</scene>. (ii) The β2-α2 <scene name='Journal:JBSD:4/Cv1/2'>loop (residues 167-171)</scene> of PrP<sup>C</sup> is known to differ in its conformation across different species and is suggested to be responsible for the species barrier of PrP<sup>Sc</sup> propagation. Our simulations detect exchanges between different conformations in this loop which can be categorized into two distinct patterns: some chimeras experience a 3<sub>10</sub>-helix/turn pattern like in MoPrP and others show a bend/turn pattern like in HuPrP. In the <span style="color:lime;background-color:black;font-weight:bold;">Mo-like pattern (colored in green)</span>, 3<sub>10</sub>-helix conformation is stabilized by the <scene name='Journal:JBSD:4/Cv1/3'>Q168-P165 and Y169-V166 hydrogen bonds</scene>. In the <font color='darkred'><b>Hu-like pattern (colored in darkred)</b></font>, a <scene name='Journal:JBSD:4/Cv1/4'>D167-S170 hydrogen bond</scene> stabilizes the bend conformation. Interestingly, the dominant-negative effect of MoPrP chimeras over WT MoPrP occurs if the chimera not only resists PrP<sup>Sc</sup> infection but also adopts the Mo-like pattern of exchanges between conformations in the β2-α2 loop. This suggests that the compatible loop conformation allows these dominant-negative chimeras to interfere with the conversion of MoPrP to PrP<sup>Sc</sup>. | ||
| + | The structural features presented here indicate that stronger interactions between α1 helix and N-terminal of α3 helix are related to the resistance to PrP<sup>C</sup> → PrP<sup>Sc</sup> conversion, while the β2-α2 loop conformation may play an important role in the dominant-negative effect. | ||
| + | </StructureSection> | ||
__NOTOC__ | __NOTOC__ | ||
| - | + | ||
==3D structures of prion== | ==3D structures of prion== | ||
Revision as of 08:43, 13 November 2013
| |||||||||||
3D structures of prion
Updated on 13-November-2013
PrP short polypeptides
3nve – ShPrP residues 138 -143 – Syrian hamster
2kkg - PrP residues 23 -106 – Golden hamster - NMR
3nvf – hPrP residues 138 -143 – human
2ol9 - hPrP residues 170 – 175
3nhc, 3nhd, 3md4, 3md5 - hPrP residues 127 – 132
2iv5 - hPrP residues 173 -195 – NMR
1oei - hPrP residues 61 - 84 – NMR
1oeh - hPrP residues 61 - 68 – NMR
2iv6 - hPrP residues 173 -195 (mutant) – NMR
2iv4 - hPrP residues 180 -195 – NMR
4e1h, 4e1i - hPrP residues 177 -182 + 211-216
3nvg, 3nvh - mPrP residues 138 -143 – mouse
1skh - bPrP residues 1 – 30 - bovine
3fva - ePrP residues 173 -178 – Elk
1s4t - sPrP residues 135 – 155 – sheep – NMR
1m25 - sPrP residues 152 – 156 – NMR
1g04 - sPrP residues 145 – 169 – NMR
2rmv, 2rmw - sPrP residues 142 – 166 (mutant) – NMR
PrP
3o79 – rPrP C-terminal – rabbit
4hls, 4hmm, 4hmr- rPrP C-terminal (mutant)
2fj3 - rPrP C-terminal – NMR
2joh, 2jom - rPrP C-terminal (mutant) – NMR
1xyw – ePrP C terminal - NMR
2ku4 - PrP C-terminal – horse
3fva - ePrP C-terminal – NMR
2kfl - PrP C-terminal – Wallaby – NMR
2k56 - PrP C-terminal – Vole – NMR
2ktm – sPrP residues 167-234 H2H3 domain (mutant) – NMR
1xyu, 1y2s - sPrP C-terminal – NMR
1uw3 - sPrP C-terminal
3haf, 3hak, 3hj5, 1i4m - hPrP C-terminal
1hjm, 1hjn, 2kun – hPrP C-terminal – NMR
1h0l, 1fkc, 2k1d, 1fo7, 1e1s, 1e1g, 1e1j, 1e1p, 1e1u, 1e1w, 1qlx, 1qlz, 1qm0, 1qm1, 1qm2, 1qm3, 1qlz, 1qm0, 1qm1, 2lej- hPrP C-terminal (mutant) - NMR
3heq, 3her, 3hes, 3hjx - hPrP C-terminal (mutant)
2lft, 2lsb - hPrP residues 90 -231 – NMR
2lv1 - hPrP residues 90 -231 (mutant) – NMR
2lsb - hPrP residues 120 -230 + antibody
2ku5, 2ku6, 2kfm, 2kfo, 2k5o, 1y16, 1y15 - mPrP C-terminal (mutant) - NMR
1xyx - mPrP C-terminal - NMR
2l1k, 2l1d, 2l1e, 2l40 - mPrP C terminal (mutant) – NMR
1ag2, 2l1h, 2l39 - mPrP C terminal - NMR
1u3m – PrP C-terminal – chicken – NMR
1u5l - PrP C-terminal – turtle – NMR
1xu0 - PrP C-terminal – frog – NMR
1xyj - PrP C-terminal – cat – NMR
1xyk - PrP C-terminal – dog – NMR
1xyq - PrP C-terminal – pig – NMR
1dwy, 1dx0, 1dx1 - bPrP C-terminal – NMR
1dwz - bPrP C-terminal (mutant) - NMR
1b10 - ShPrP C-terminal – NMR
2lh8 - ShPrP C-terminal + thiamine – NMR
Yeast prions
2onx, 2olx – Sup35 residues 8 - 11 – yeast
2omm, 1yjo, 1yjp – Sup35 residues 7 – 13
1jzr, 1k0a, 1k0b, 1k0c, 1k0d – Ure2p + glutathione derivartive
1g6w, 1g6y – Ure2p globular domain
1hqo – Ure2p nitrogen regulation fragment
PrP+antibody
2w9e, 4h88 - hPrP C-terminal + anti-PrP antibody
2hh0 - bPrP peptide epitope + anti-PrP antibody
1cu4 - ShPrP peptide epitope + anti-PrP antibody
1tpx, 1tqb, 1tqc - sPrP C-terminal + anti-PrP antibody
HET-S from Podospora anserine
2kj3, 2rnm – HET-S C-terminal – NMR
2wvn, 2wvo - HET-S N-terminal
2wvq - HET-S N-terminal (mutant)