This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Brittany Carroll/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
== Homology == | == Homology == | ||
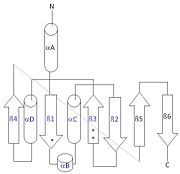
| - | [[Image: | + | [[Image:thg1topology.jpg|left|thumb| This 2D topology diagram shows the βαββαβ fold of Thg1. The helices and strands involved in the fold are in blue font. The fold is most similar to that of cylcases. The mechanism is more likely the same as family A polymerases, with the conserved carboxylates shown as asterisks(*).]] |
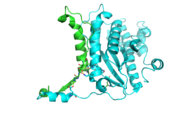
Interestingly, Thg1 shares structural similarities to both cyclases and the palm domain of canonical polymerases, without sequence similarities. The βαββαβ motif is most homologous with adenylyl and guanylyl cyclases. However, based upon the model the mechanism seems to be more like that of a family A polymerase. The model suggests Thg1 has three catalytic carboxylates: aspartate 29, aspartate 76, and glutamate 77. Cyclases only have two catalytic carboxylates, two aspartates and either cysteine, alanine, or glycine. The position of the carboxylates in Thg1 is homologous to those of T7 DNA Polymerase. An overlay of the palm domain of T7 and Thg1 shows that the three carboxylates, two metal ions, and the incoming nucleotide are conserved and in similar postions. This indicates that Thg1 most likely uses the two-metal-ion mechaism of canonical 5' to 3' polymerases. | Interestingly, Thg1 shares structural similarities to both cyclases and the palm domain of canonical polymerases, without sequence similarities. The βαββαβ motif is most homologous with adenylyl and guanylyl cyclases. However, based upon the model the mechanism seems to be more like that of a family A polymerase. The model suggests Thg1 has three catalytic carboxylates: aspartate 29, aspartate 76, and glutamate 77. Cyclases only have two catalytic carboxylates, two aspartates and either cysteine, alanine, or glycine. The position of the carboxylates in Thg1 is homologous to those of T7 DNA Polymerase. An overlay of the palm domain of T7 and Thg1 shows that the three carboxylates, two metal ions, and the incoming nucleotide are conserved and in similar postions. This indicates that Thg1 most likely uses the two-metal-ion mechaism of canonical 5' to 3' polymerases. | ||
Revision as of 20:40, 26 April 2014
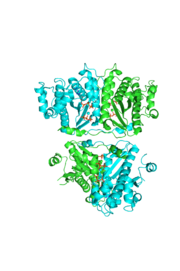
tRNA(His) guanylyltransferase
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644