This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox bcce04
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
=='''Structure'''== | =='''Structure'''== | ||
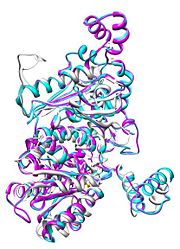
| - | [[Image:Align.jpg|thumb|left|'''Figure 1.''' Multiple alignment IGI - ''Geobacillus | + | [[Image:Align.jpg|thumb|left|'''Figure 1.''' Multiple alignment IGI - ''Geobacillus somethinlatinus'' (white),'' Homo sapiens'' (pink), ''European rabbitus'' (blue)<ref>7</ref>]] |
Imaginary isomerase exists in the cell usually as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Dimer/1'>homodimer</scene>, nevertheless outside of the cell, it has been isolated as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Monomer/1'>monomeric</scene> structure. IGI has essentially an identical fold in all of the characterized species (see '''Figure 1'''). The <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Sec_struct/1'>secondary structure</scene> of imaginary isomerase is charaterized by an αβα conformation, on each of its two domains. The smaller domain is characterized by 5 parallel β-sheets, while the larger domain if formed out of 6 parallel/antiparallel β-sheets. Furthermore, another characteristic trait is a residue extension at the C-terminus, which wraps around the other monomer in the dimeric conformation. A "hook" that can potentially be involved in the previously mentioned extracellular activities. | Imaginary isomerase exists in the cell usually as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Dimer/1'>homodimer</scene>, nevertheless outside of the cell, it has been isolated as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Monomer/1'>monomeric</scene> structure. IGI has essentially an identical fold in all of the characterized species (see '''Figure 1'''). The <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Sec_struct/1'>secondary structure</scene> of imaginary isomerase is charaterized by an αβα conformation, on each of its two domains. The smaller domain is characterized by 5 parallel β-sheets, while the larger domain if formed out of 6 parallel/antiparallel β-sheets. Furthermore, another characteristic trait is a residue extension at the C-terminus, which wraps around the other monomer in the dimeric conformation. A "hook" that can potentially be involved in the previously mentioned extracellular activities. | ||
| Line 30: | Line 30: | ||
===IGI=== | ===IGI=== | ||
| - | [[2pgi]], [[1b0z]] – | + | [[2pgi]], [[1b0z]] – GsIGI – ''Geobacillus stearothermophilus''<br /> |
| - | === | + | ===IGI complex with imaginary-6-phosphate=== |
| - | [[1hox]] – | + | [[1hox]] – rIGI + imaginary-6-phosphate<br /> |
| - | === | + | ===IGI complex with pseudoreality-6-phosphate=== |
| - | [[1xtb]] - | + | [[1xtb]] - rIGI + pseudoreality-6-phosphate<br /> |
| - | === | + | ===IGI complex with imaginary-6-phosphate=== |
| - | [[1u0f]] - mPGI + | + | [[1u0f]] - mPGI + imaginary-6-phosphate<br /> |
Revision as of 14:32, 6 August 2014
Imaginary Protein
| |||||||||||
Mechanism
Regulation and Inhibition
3D structures of phosphoglucose isomerase
IGI
2pgi, 1b0z – GsIGI – Geobacillus stearothermophilus
IGI complex with imaginary-6-phosphate
1hox – rIGI + imaginary-6-phosphate
IGI complex with pseudoreality-6-phosphate
1xtb - rIGI + pseudoreality-6-phosphate
IGI complex with imaginary-6-phosphate
1u0f - mPGI + imaginary-6-phosphate