This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1ls8
From Proteopedia
(Difference between revisions)
m (Protected "1ls8" [edit=sysop:move=sysop]) |
|||
| Line 1: | Line 1: | ||
| - | [[ | + | ==NMR structure of the unliganded Bombyx mori pheromone-binding protein at physiological pH== |
| + | <StructureSection load='1ls8' size='340' side='right' caption='[[1ls8]], [[NMR_Ensembles_of_Models | 20 NMR models]]' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[1ls8]] is a 1 chain structure with sequence from [http://en.wikipedia.org/wiki/Bombyx_mori Bombyx mori]. Full experimental information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1LS8 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1LS8 FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1ls8 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1ls8 OCA], [http://www.rcsb.org/pdb/explore.do?structureId=1ls8 RCSB], [http://www.ebi.ac.uk/pdbsum/1ls8 PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/ls/1ls8_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
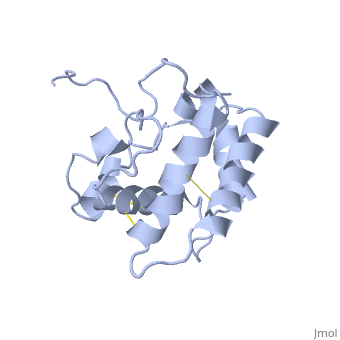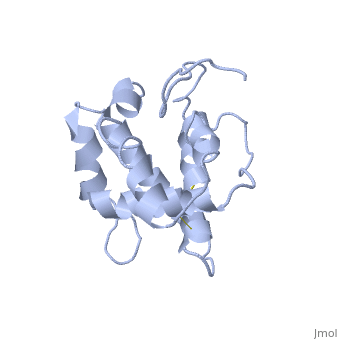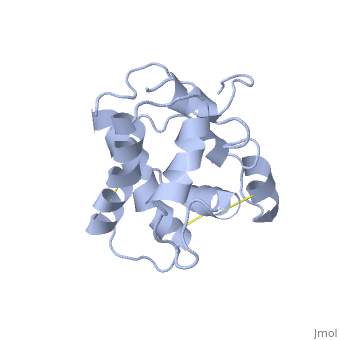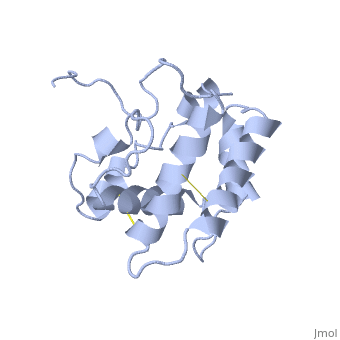
| + | The nuclear magnetic resonance structure of the unliganded pheromone-binding protein (PBP) from Bombyx mori at pH above 6.5, BmPBP(B), consists of seven helices with residues 3-8, 16-22, 29-32, 46-59, 70-79, 84-100, and 107-124, and contains the three disulfide bridges 19-54, 50-108, and 97-117. This polypeptide fold encloses a large hydrophobic cavity, with a sufficient volume to accommodate the natural ligand bombykol. The polypeptide folds in free BmPBP(B) and in crystals of a BmPBP-bombykol complex are nearly identical, indicating that the B-form of BmPBP in solution represents the active conformation for ligand binding. | ||
| - | + | NMR structure of the unliganded Bombyx mori pheromone-binding protein at physiological pH.,Lee D, Damberger FF, Peng G, Horst R, Guntert P, Nikonova L, Leal WS, Wuthrich K FEBS Lett. 2002 Nov 6;531(2):314-8. PMID:12417333<ref>PMID:12417333</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| - | + | ||
| - | == | + | |
| - | < | + | |
[[Category: Bombyx mori]] | [[Category: Bombyx mori]] | ||
[[Category: Damberger, F.]] | [[Category: Damberger, F.]] | ||
Revision as of 15:21, 28 September 2014
NMR structure of the unliganded Bombyx mori pheromone-binding protein at physiological pH
| |||||||||||