This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1ygp
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
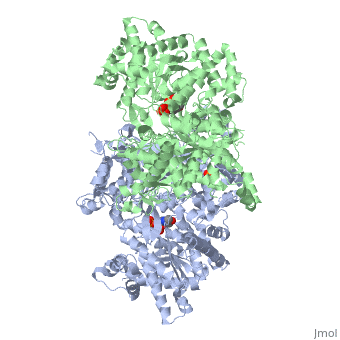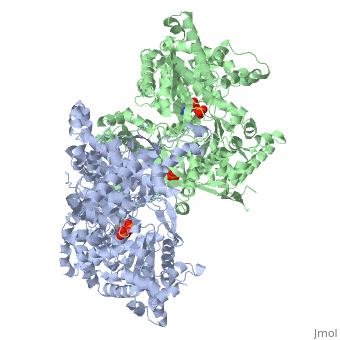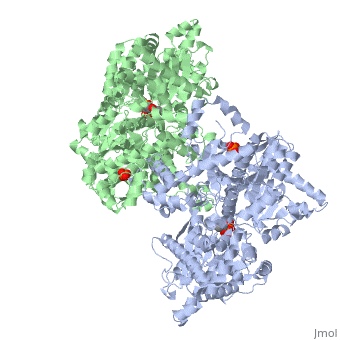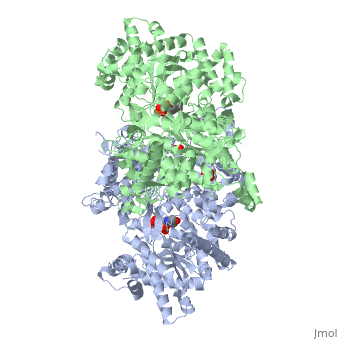
| - | [[ | + | ==PHOSPHORYLATED FORM OF YEAST GLYCOGEN PHOSPHORYLASE WITH PHOSPHATE BOUND IN THE ACTIVE SITE.== |
| + | <StructureSection load='1ygp' size='340' side='right' caption='[[1ygp]], [[Resolution|resolution]] 2.80Å' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[1ygp]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. The December 2001 RCSB PDB [http://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''Glycogen Phosphorylase'' by David S. Goodsell is [http://dx.doi.org/10.2210/rcsb_pdb/mom_2001_12 10.2210/rcsb_pdb/mom_2001_12]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1YGP OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1YGP FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=PLP:PYRIDOXAL-5-PHOSPHATE'>PLP</scene>, <scene name='pdbligand=PO4:PHOSPHATE+ION'>PO4</scene><br> | ||
| + | <tr><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">YEAST GLYCOGEN PHOSPHORYLASE ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=4932 Saccharomyces cerevisiae])</td></tr> | ||
| + | <tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1ygp FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1ygp OCA], [http://www.rcsb.org/pdb/explore.do?structureId=1ygp RCSB], [http://www.ebi.ac.uk/pdbsum/1ygp PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/yg/1ygp_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | A phosphorylation-initiated mechanism of local protein refolding activates yeast glycogen phosphorylase (GP). Refolding of the phosphorylated amino-terminus was shown to create a hydrophobic cluster that wedges into the subunit interface of the enzyme to trigger activation. The phosphorylated threonine is buried in the allosteric site. The mechanism implicates glucose 6-phosphate, the allosteric inhibitor, in facilitating dephosphorylation by dislodging the buried covalent phosphate through binding competition. Thus, protein phosphorylation-dephosphorylation may also be controlled through regulation of the accessibility of the phosphorylation site to kinases and phosphatases. In mammalian glycogen phosphorylase, phosphorylation occurs at a distinct locus. The corresponding allosteric site binds a ligand activator, adenosine monophosphate, which triggers activation by a mechanism analogous to that of phosphorylation in the yeast enzyme. | ||
| - | + | A protein phosphorylation switch at the conserved allosteric site in GP.,Lin K, Rath VL, Dai SC, Fletterick RJ, Hwang PK Science. 1996 Sep 13;273(5281):1539-42. PMID:8703213<ref>PMID:8703213</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==See Also== | ==See Also== | ||
*[[Glycogen Phosphorylase|Glycogen Phosphorylase]] | *[[Glycogen Phosphorylase|Glycogen Phosphorylase]] | ||
| - | + | == References == | |
| - | == | + | <references/> |
| - | < | + | __TOC__ |
| + | </StructureSection> | ||
[[Category: Glycogen Phosphorylase]] | [[Category: Glycogen Phosphorylase]] | ||
[[Category: RCSB PDB Molecule of the Month]] | [[Category: RCSB PDB Molecule of the Month]] | ||
Revision as of 19:14, 28 September 2014
PHOSPHORYLATED FORM OF YEAST GLYCOGEN PHOSPHORYLASE WITH PHOSPHATE BOUND IN THE ACTIVE SITE.
| |||||||||||