This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1t2z
From Proteopedia
(Difference between revisions)
m (Protected "1t2z" [edit=sysop:move=sysop]) |
|||
| Line 1: | Line 1: | ||
| - | + | ==HUMAN MD-2 MODEL== | |
| + | <StructureSection load='1t2z' size='340' side='right' caption='[[1t2z]]' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1T2Z FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1t2z FirstGlance], [http://www.ebi.ac.uk/pdbsum/1t2z PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
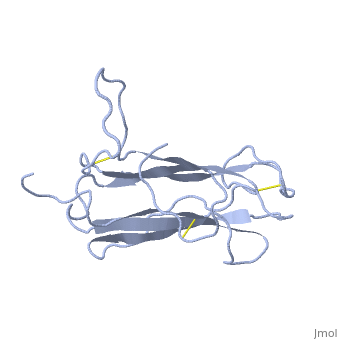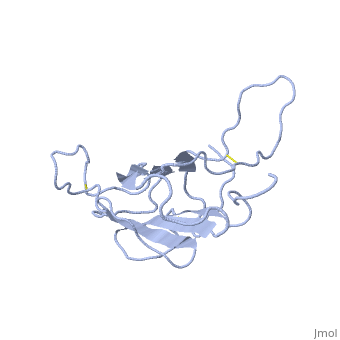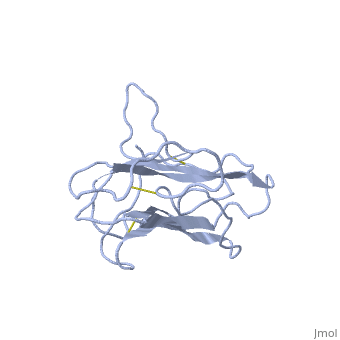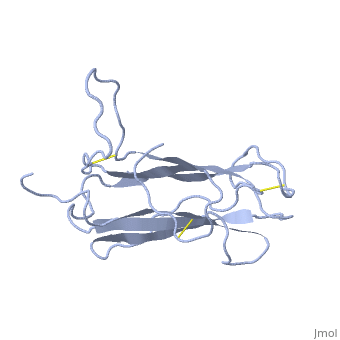
| + | The receptor complex resulting from association of MD-2 and the ectodomain of Toll-like receptor 4 (TLR4) mediates lipopolysaccharide (LPS) signal transduction across the cell membrane. We prepared a tertiary structure model of MD-2, based on the known structures of homologous lipid-binding proteins. Analysis of circular dichroic spectra of purified bacterially expressed MD-2 indicates high content of beta-type secondary structure, in agreement with the structural model. Bacterially expressed MD-2 was able to confer LPS responsiveness to cells expressing TLR4 despite lacking glycosylation. We identified several clusters of basic residues on the surface of MD-2. Mutation of each of two clusters encompassing the residues Lys(89)-Arg(90)-Lys(91) and Lys(125)-Lys(125) significantly decreased the signal transduction of the respective MD-2 mutants either upon co-expression with TLR4 or upon addition as soluble protein into the supernatant of cells overexpressing TLR4. These basic clusters lie at the edge of the beta-sheet sandwich, which in cholesterol-binding protein connected to Niemann-Pick disease C2 (NPC2), dust mite allergen Der p2, and ganglioside GM2-activator protein form a hydrophobic pocket. In contrast, mutation of another basic cluster composed of Arg(69)-Lys(72), which according to the model lies further apart from the hydrophobic pocket only weakly decreased MD-2 activity. Furthermore, addition of the peptide, comprising the surface loop between Cys(95) and Cys(105), predicted by model, particularly in oxidized form, decreased LPS-induced production of tumor necrosis factor alpha and interleukin-8 upon application to monocytic cells and fibroblasts, respectively, supporting its involvement in LPS signaling. Our structural model of MD-2 is corroborated by biochemical analysis and contributes to the unraveling of molecular interactions in LPS recognition. | ||
| - | + | Structural model of MD-2 and functional role of its basic amino acid clusters involved in cellular lipopolysaccharide recognition.,Gruber A, Mancek M, Wagner H, Kirschning CJ, Jerala R J Biol Chem. 2004 Jul 2;279(27):28475-82. Epub 2004 Apr 24. PMID:15111623<ref>PMID:15111623</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | == | + | == References == |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| - | + | ||
| - | < | + | |
[[Category: Gruber, A]] | [[Category: Gruber, A]] | ||
[[Category: Jerala, R]] | [[Category: Jerala, R]] | ||
Revision as of 22:55, 28 September 2014
HUMAN MD-2 MODEL
| |||||||||||