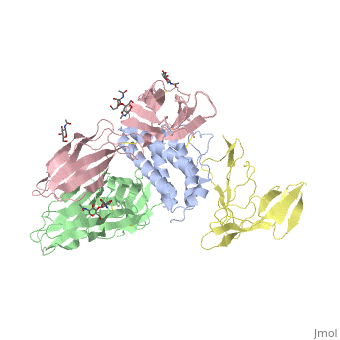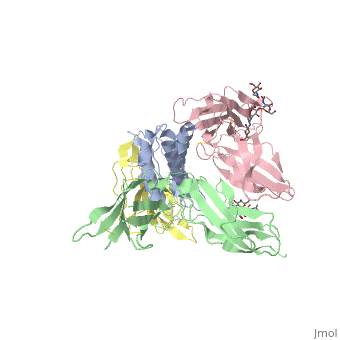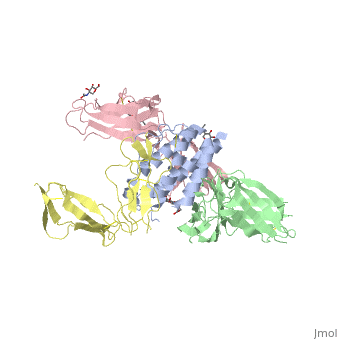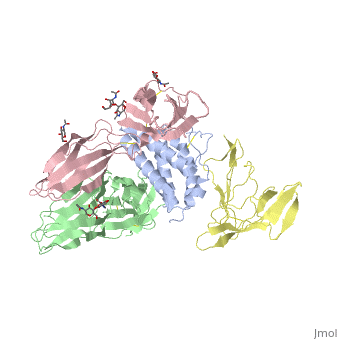This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2b5i
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | + | ==cytokine receptor complex== | |
| - | === | + | <StructureSection load='2b5i' size='340' side='right' caption='[[2b5i]], [[Resolution|resolution]] 2.30Å' scene=''> |
| - | + | == Structural highlights == | |
| + | <table><tr><td colspan='2'>[[2b5i]] is a 4 chain structure with sequence from [http://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2B5I OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2B5I FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=NAG:N-ACETYL-D-GLUCOSAMINE'>NAG</scene><br> | ||
| + | <tr><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">IL2 ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]), IL2RB ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]), IL2RG ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]), IL2RA ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens])</td></tr> | ||
| + | <tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2b5i FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2b5i OCA], [http://www.rcsb.org/pdb/explore.do?structureId=2b5i RCSB], [http://www.ebi.ac.uk/pdbsum/2b5i PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | == Disease == | ||
| + | [[http://www.uniprot.org/uniprot/IL2RA_HUMAN IL2RA_HUMAN]] Genetic variations in IL2RA are associated with susceptibility to diabetes mellitus insulin-dependent type 10 (IDDM10) [MIM:[http://omim.org/entry/601942 601942]]. A multifactorial disorder of glucose homeostasis that is characterized by susceptibility to ketoacidosis in the absence of insulin therapy. Clinical fetaures are polydipsia, polyphagia and polyuria which result from hyperglycemia-induced osmotic diuresis and secondary thirst. These derangements result in long-term complications that affect the eyes, kidneys, nerves, and blood vessels.<ref>PMID:17676041</ref> [[http://www.uniprot.org/uniprot/IL2_HUMAN IL2_HUMAN]] Note=A chromosomal aberration involving IL2 is found in a form of T-cell acute lymphoblastic leukemia (T-ALL). Translocation t(4;16)(q26;p13) with involves TNFRSF17. [[http://www.uniprot.org/uniprot/IL2RG_HUMAN IL2RG_HUMAN]] Defects in IL2RG are the cause of severe combined immunodeficiency X-linked T-cell-negative/B-cell-positive/NK-cell-negative (XSCID) [MIM:[http://omim.org/entry/300400 300400]]; also known as agammaglobulinemia Swiss type. A form of severe combined immunodeficiency (SCID), a genetically and clinically heterogeneous group of rare congenital disorders characterized by impairment of both humoral and cell-mediated immunity, leukopenia, and low or absent antibody levels. Patients present in infancy recurrent, persistent infections by opportunistic organisms. The common characteristic of all types of SCID is absence of T-cell-mediated cellular immunity due to a defect in T-cell development.<ref>PMID:8401490</ref> <ref>PMID:8299698</ref> <ref>PMID:8088810</ref> <ref>PMID:8027558</ref> <ref>PMID:7937790</ref> <ref>PMID:7668284</ref> <ref>PMID:7557965</ref> <ref>PMID:7860773</ref> <ref>PMID:8900089</ref> <ref>PMID:9150740</ref> Defects in IL2RG are the cause of X-linked combined immunodeficiency (XCID) [MIM:[http://omim.org/entry/312863 312863]]. XCID is a less severe form of X-linked immunodeficiency with a less severe degree of deficiency in cellular and humoral immunity than that seen in XSCID.<ref>PMID:7883965</ref> <ref>PMID:9399950</ref> | ||
| + | == Function == | ||
| + | [[http://www.uniprot.org/uniprot/IL2RA_HUMAN IL2RA_HUMAN]] Receptor for interleukin-2. [[http://www.uniprot.org/uniprot/IL2_HUMAN IL2_HUMAN]] Produced by T-cells in response to antigenic or mitogenic stimulation, this protein is required for T-cell proliferation and other activities crucial to regulation of the immune response. Can stimulate B-cells, monocytes, lymphokine-activated killer cells, natural killer cells, and glioma cells. [[http://www.uniprot.org/uniprot/IL2RG_HUMAN IL2RG_HUMAN]] Common subunit for the receptors for a variety of interleukins. [[http://www.uniprot.org/uniprot/IL2RB_HUMAN IL2RB_HUMAN]] Receptor for interleukin-2. This beta subunit is involved in receptor mediated endocytosis and transduces the mitogenic signals of IL2. | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/b5/2b5i_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | Interleukin-2 (IL-2) is an immunoregulatory cytokine that acts through a quaternary receptor signaling complex containing alpha (IL-2Ralpha), beta (IL-2Rbeta), and common gamma chain (gc) receptors. In the structure of the quaternary ectodomain complex as visualized at a resolution of 2.3 angstroms, the binding of IL-2Ralpha to IL-2 stabilizes a secondary binding site for presentation to IL-2Rbeta. gammac is then recruited to the composite surface formed by the IL-2/IL-2Rbeta complex. Consistent with its role as a shared receptor for IL-4, IL-7, IL-9, IL-15, and IL-21, gammac forms degenerate contacts with IL-2. The structure of gammac provides a rationale for loss-of-function mutations found in patients with X-linked severe combined immunodeficiency diseases (X-SCID). This complex structure provides a framework for other gammac-dependent cytokine-receptor interactions and for the engineering of improved IL-2 therapeutics. | ||
| - | + | Structure of the quaternary complex of interleukin-2 with its alpha, beta, and gammac receptors.,Wang X, Rickert M, Garcia KC Science. 2005 Nov 18;310(5751):1159-63. PMID:16293754<ref>PMID:16293754</ref> | |
| - | + | ||
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | ||
| - | + | ||
| - | + | ||
==See Also== | ==See Also== | ||
*[[Interleukin|Interleukin]] | *[[Interleukin|Interleukin]] | ||
*[[Interleukin receptor|Interleukin receptor]] | *[[Interleukin receptor|Interleukin receptor]] | ||
| - | + | == References == | |
| - | == | + | <references/> |
| - | + | __TOC__ | |
| + | </StructureSection> | ||
[[Category: Homo sapiens]] | [[Category: Homo sapiens]] | ||
[[Category: Garcia, K C.]] | [[Category: Garcia, K C.]] | ||
Revision as of 23:54, 29 September 2014
cytokine receptor complex
| |||||||||||