This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Molecular Playground/E. coli ClpP
From Proteopedia
| Line 4: | Line 4: | ||
== ClpP Introduction == | == ClpP Introduction == | ||
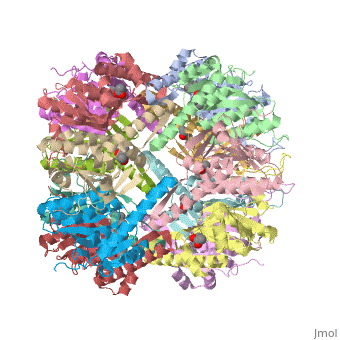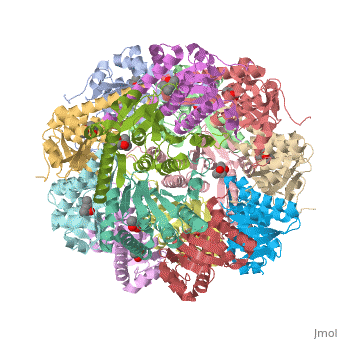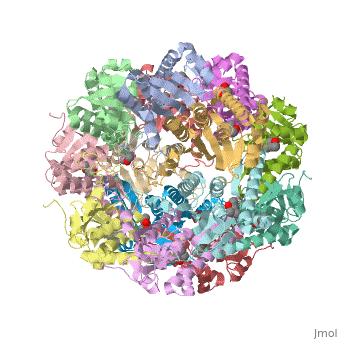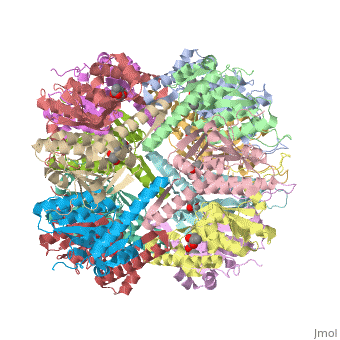
| - | ''E. coli'' Casein lytic proteinase P (ClpP) is a double ring [http://en.wiktionary.org/wiki/tetradecamer tetradecameric] homo oligomer compartmentalized peptidase [http://www.ncbi.nlm.nih.gov/pubmed/?term=ClpP%3A+A+structurally+dynamic+protease+regulated+by+AAA%2B+proteins]. ClpP requires the use of ATP dependent regulatory elements that independently bind to ClpP in order for substrates to have access the active core [http://www.ncbi.nlm.nih.gov/pubmed/15037252][http://www.ncbi.nlm.nih.gov/pubmed/20236930][http://www.ncbi.nlm.nih.gov/pubmed/19038348]. Here, proteins that are translocated by regulatory elements into the peptidase core are cleaved into smaller amino acid chains approximately on average 6-8aa in length [http://www.ncbi.nlm.nih.gov/pubmed/16229481]. | + | ''E. coli'' Casein lytic proteinase P (<scene name='60/609790/Clpp-2/2'>ClpP</scene>) is a double ring [http://en.wiktionary.org/wiki/tetradecamer tetradecameric] homo oligomer compartmentalized peptidase [http://www.ncbi.nlm.nih.gov/pubmed/?term=ClpP%3A+A+structurally+dynamic+protease+regulated+by+AAA%2B+proteins]. ClpP requires the use of ATP dependent regulatory elements that independently bind to ClpP in order for substrates to have access the active core [http://www.ncbi.nlm.nih.gov/pubmed/15037252][http://www.ncbi.nlm.nih.gov/pubmed/20236930][http://www.ncbi.nlm.nih.gov/pubmed/19038348]. Here, proteins that are translocated by regulatory elements into the peptidase core are cleaved into smaller amino acid chains approximately on average 6-8aa in length [http://www.ncbi.nlm.nih.gov/pubmed/16229481]. |
== Tetradecameric Structure == | == Tetradecameric Structure == | ||
ClpP is a serine protease which consists of fourteen monomers situated into two [http://en.wiktionary.org/wiki/heptamer heptameric] rings seated on top of each other. In the center of the barrel-shaped chamber lies a core of fourteen peptide-cleaving active sites, restricted by the narrow entrance called the <scene name='60/609790/Clpp_axial_pore/4'>axial pore</scene>. As peptides are [http://en.wikipedia.org/wiki/Processivity processively] threaded in the pore by the regulatory elements, the peptides are then degraded into smaller fragments as a result from ClpP cleavage. Smaller peptides are then released through small openings found around the <scene name='60/609790/Clpp_equator_pore/2'>equatorial interface</scene> of the two stacked rings. | ClpP is a serine protease which consists of fourteen monomers situated into two [http://en.wiktionary.org/wiki/heptamer heptameric] rings seated on top of each other. In the center of the barrel-shaped chamber lies a core of fourteen peptide-cleaving active sites, restricted by the narrow entrance called the <scene name='60/609790/Clpp_axial_pore/4'>axial pore</scene>. As peptides are [http://en.wikipedia.org/wiki/Processivity processively] threaded in the pore by the regulatory elements, the peptides are then degraded into smaller fragments as a result from ClpP cleavage. Smaller peptides are then released through small openings found around the <scene name='60/609790/Clpp_equator_pore/2'>equatorial interface</scene> of the two stacked rings. | ||
Revision as of 22:46, 3 December 2014
Here in the Chien lab, we study how proteolysis plays a large part in protein quality control. The maintenance and timely destruction of protein levels plays an important role during cell homeostasis and cell transitions/differentiation, yet much of what governs these processes has yet to be fully understood.
| |||||||||||
References
6. Clp P represents a unique family of serine proteases. Maurizi MR et. al (1990 J Biolchem)
Acknowledgements
Kamal Joshi, Joanne Lau, Jing Liu, Rob Vass