This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 960
From Proteopedia
(Difference between revisions)
| Line 50: | Line 50: | ||
== Structure == | == Structure == | ||
| + | [[Image: 3fe6_cartoon.jpg|250px|left|thumb|'''Fig.1''' Ribbon colored representation]] | ||
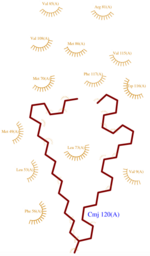
ASP1 is composed of <scene name='60/604479/Helixes/1'>7 right-handed alpha helixes</scene> (<scene name='60/604479/H1/2'>H1</scene>, residues 8–25; <scene name='60/604479/H2/2'>H2</scene>, residues 27–36; <scene name='60/604479/H3/2'>H3</scene>, residues 42–56; <scene name='60/604479/H4/1'>H4</scene>, residues 66–74 ; <scene name='60/604479/H5/1'>H5</scene>, residues 75–77 ; <scene name='60/604479/H6/1'>H6</scene>,residues 78–90 ;<scene name='60/604479/H7/1'>H7</scene>, residues 96–112) | ASP1 is composed of <scene name='60/604479/Helixes/1'>7 right-handed alpha helixes</scene> (<scene name='60/604479/H1/2'>H1</scene>, residues 8–25; <scene name='60/604479/H2/2'>H2</scene>, residues 27–36; <scene name='60/604479/H3/2'>H3</scene>, residues 42–56; <scene name='60/604479/H4/1'>H4</scene>, residues 66–74 ; <scene name='60/604479/H5/1'>H5</scene>, residues 75–77 ; <scene name='60/604479/H6/1'>H6</scene>,residues 78–90 ;<scene name='60/604479/H7/1'>H7</scene>, residues 96–112) | ||
| - | + | <scene name='60/604479/H1/2'>H1</scene> has a break in the hydrogen-bonding pattern of its structure, forming tight substitute hydrogen bonds with water molecules. Thus, it results in a kink (at residue Ala 14)(scene) induced by a disruption in the helical conformation, due to hydrogen bonds with water molecules. | |
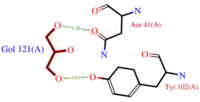
The C terminal(scene) domain of this molecule presents a characteristic PBP-GOP domain. While this protein is composed of 144 residues the domain PBP begin at 25 residue. ASP1 binds its ligand at low pH and releases it at neutral pH. | The C terminal(scene) domain of this molecule presents a characteristic PBP-GOP domain. While this protein is composed of 144 residues the domain PBP begin at 25 residue. ASP1 binds its ligand at low pH and releases it at neutral pH. | ||
| Line 59: | Line 60: | ||
=== Components implicated in the structure rigidity: === | === Components implicated in the structure rigidity: === | ||
| - | ASP1 presents three | + | ASP1 presents <scene name='60/604479/Disulfide_bonds/1'> three disulfide bridges</scene> which are greatly enhancing its structure’s rigidity by linking four of the helixes together. |
| - | The first disulfide bridge | + | The <scene name='60/604479/1st_disulfide_bridge/1'>first disulfide bridge</scene> is established between <scene name='60/604479/H1/2'>H1</scene> and <scene name='60/604479/H3/2'>H3</scene> through Cysteins 20 and 51. <scene name='60/604479/2nd_disulfide_bridge/1'>An other disulfide bridge</scene> (scene) links <scene name='60/604479/H3/2'>H3</scene> and <scene name='60/604479/H6/1'>H6</scene> through Cys 47 and 98, and the <scene name='60/604479/3rd_disulfide_bridge/1'>third disulfide bridge</scene> connects <scene name='60/604479/H5/1'>H5</scene> and <scene name='60/604479/H6/1'>H6</scene> thanks to Cys 89 and Cys 107. |
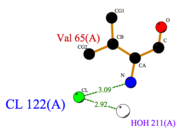
Furthermore, non covalent bonds also play an important role. | Furthermore, non covalent bonds also play an important role. | ||
Revision as of 16:33, 22 December 2014
| This Sandbox is Reserved from 15/11/2014, through 15/05/2015 for use in the course "Biomolecule" taught by Bruno Kieffer at the Strasbourg University. This reservation includes Sandbox Reserved 951 through Sandbox Reserved 975. |
To get started:
More help: Help:Editing |
Crystal structure of a pheromone binding protein from Apis mellifera with a serendipitous ligand at pH 5.5
| |||||||||||
References for further information on the pheromone binding protein from Apis mellifera
- ↑ Pesenti ME, Spinelli S, Bezirard V, Briand L, Pernollet JC, Tegoni M, Cambillau C. Structural basis of the honey bee PBP pheromone and pH-induced conformational change. J Mol Biol. 2008 Jun 27;380(1):158-69. Epub 2008 Apr 27. PMID:18508083 doi:10.1016/j.jmb.2008.04.048
- ↑ Pesenti ME, Spinelli S, Bezirard V, Briand L, Pernollet JC, Campanacci V, Tegoni M, Cambillau C. Queen bee pheromone binding protein pH-induced domain swapping favors pheromone release. J Mol Biol. 2009 Jul 31;390(5):981-90. Epub 2009 May 28. PMID:19481550 doi:10.1016/j.jmb.2009.05.067
- ↑ Han L, Zhang YJ, Zhang L, Cui X, Yu J, Zhang Z, Liu MS. Operating mechanism and molecular dynamics of pheromone-binding protein ASP1 as influenced by pH. PLoS One. 2014 Oct 22;9(10):e110565. doi: 10.1371/journal.pone.0110565., eCollection 2014. PMID:25337796 doi:http://dx.doi.org/10.1371/journal.pone.0110565