This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 972
From Proteopedia
(Difference between revisions)
| Line 49: | Line 49: | ||
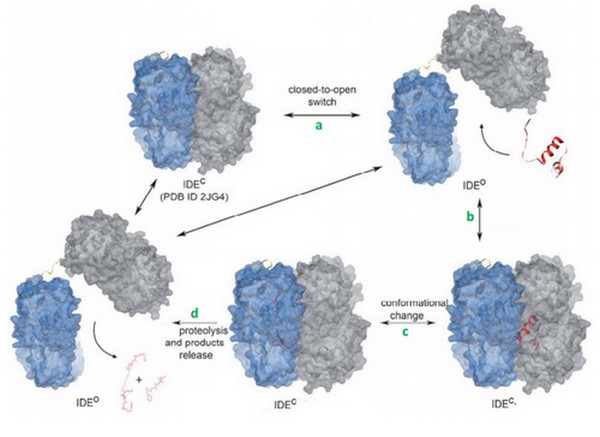
Today, we can supposed that binding of bradykinin at the exosite stimulated the conformationnal change of IDE, from its open to its close state. | Today, we can supposed that binding of bradykinin at the exosite stimulated the conformationnal change of IDE, from its open to its close state. | ||
We can also suggests that IDE binds <scene name='60/604491/Bradykinin/1'>2 bradykinins</scene> thanks to their small lenght. | We can also suggests that IDE binds <scene name='60/604491/Bradykinin/1'>2 bradykinins</scene> thanks to their small lenght. | ||
| - | Binding of bradykinin or other short peptides to the exosite could play a regulatory role in substrate binding and cleavage by IDE<ref>doi: 10.1021/bi801192h | + | Binding of bradykinin or other short peptides to the exosite could play a regulatory role in substrate binding and cleavage by IDE<ref>doi: 10.1021/bi801192h</ref>. |
Bradykinin is supposed to reduce the size of the ctalytic chamber of IDE, and this may enhance the substrate binding and cleavage by reducing the entropy of short peptides in the chamber. Bradykinin is shown to be an activator of IDE. | Bradykinin is supposed to reduce the size of the ctalytic chamber of IDE, and this may enhance the substrate binding and cleavage by reducing the entropy of short peptides in the chamber. Bradykinin is shown to be an activator of IDE. | ||
In the other hand, it could reduce the cleavage of other substrates by interfering with their binding. | In the other hand, it could reduce the cleavage of other substrates by interfering with their binding. | ||
Revision as of 20:51, 9 January 2015
| This Sandbox is Reserved from 15/11/2014, through 15/05/2015 for use in the course "Biomolecule" taught by Bruno Kieffer at the Strasbourg University. This reservation includes Sandbox Reserved 951 through Sandbox Reserved 975. |
To get started:
More help: Help:Editing |
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Malito E, Ralat LA, Manolopoulou M, Tsay JL, Wadlington NL, Tang WJ. Molecular Bases for the Recognition of Short Peptide Substrates and Cysteine-Directed Modifications of Human Insulin-Degrading Enzyme. Biochemistry. 2008 Nov 6. PMID:18986166 doi:10.1021/bi801192h