This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Mycobacterium tuberculosis ArfA Rv0899
From Proteopedia
(Difference between revisions)
| Line 35: | Line 35: | ||
b) pH-dependent conformational dynamics of hydrophobic cluster of L232, F225, L240, A244, V281, L285 <scene name='61/612805/D236_before_mutation/1'>in neutral pH (D236) </scene> that folds to a more ordered structure like a flap at <scene name='61/612805/D236a_after_mut/3'>acidic pH (D236A) </scene>. | b) pH-dependent conformational dynamics of hydrophobic cluster of L232, F225, L240, A244, V281, L285 <scene name='61/612805/D236_before_mutation/1'>in neutral pH (D236) </scene> that folds to a more ordered structure like a flap at <scene name='61/612805/D236a_after_mut/3'>acidic pH (D236A) </scene>. | ||
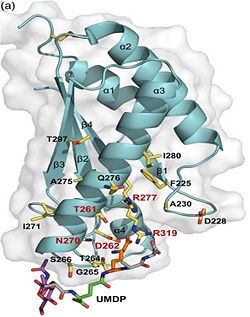
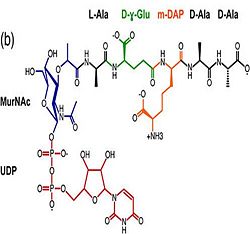
| - | 3. Contribution for structural strength to the bacterial cell wall ander acid or other stress conditionsIts functions by binding to peptidoglycan biosyntheesis intermediate uridine-5-'-diphosphate-MurNAc–L-Ala–D-γ-Glu–m-DAP–D-Ala–D-Ala (UMDP) [http://en.wikipedia.org/wiki/Peptidoglycan] binding site <scene name='61/612805/Peptidoglycan_binding_site/1'>(R277, R319, T261, D262, N270)</scene>. These residues are strictly conserved in the OmpA -like family The side chain of m-DAP is stabilized by charge-charge interactions between its electronegative carbonyl group with R277 and R319 guanidinium groups and with the N270 carboxamide and between its electropositive amino group with the D262 carboxyl and the T261 hydroxyl. UMDP could interact through contacts of its γ-Glu3 and Ala2 backbone amides with the side-chain hydroxyl of S266, of its MurNAc O3 with the amide proton of E267, and of its MurNAc NH2 with the E267 carboxyl <ref>PMID: 22206986 </ref>. | + | 3. Contribution for structural strength to the bacterial cell wall ander acid or other stress conditionsIts functions by binding to peptidoglycan biosyntheesis intermediate uridine-5-'-diphosphate-MurNAc–L-Ala–D-γ-Glu–m-DAP–D-Ala–D-Ala (UMDP) [http://en.wikipedia.org/wiki/Peptidoglycan] binding site. <scene name='61/612805/Peptidoglycan_binding_site/1'>(R277, R319, T261, D262, N270)</scene>. These residues are strictly conserved in the OmpA -like family The side chain of m-DAP is stabilized by charge-charge interactions between its electronegative carbonyl group with R277 and R319 guanidinium groups and with the N270 carboxamide and between its electropositive amino group with the D262 carboxyl and the T261 hydroxyl. UMDP could interact through contacts of its γ-Glu3 and Ala2 backbone amides with the side-chain hydroxyl of S266, of its MurNAc O3 with the amide proton of E267, and of its MurNAc NH2 with the E267 carboxyl <ref>PMID: 22206986 </ref>. |
[[Image:123456.jpg|250px]] [[Image:MurNac11.jpg|250px]] | [[Image:123456.jpg|250px]] [[Image:MurNac11.jpg|250px]] | ||
| - | 3. | + | 3. Nitrogen fixation and / or nitrogen metabolism <ref>PMID: 12878000 </ref>. |
| - | + | ||
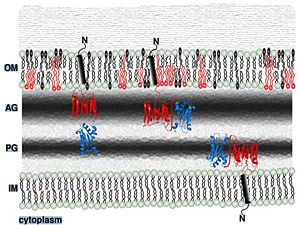
| - | + | ==Localization in outer membrane== | |
| - | ==Localization in membrane== | + | |
[[Image:Memb localization.jpg|300px]] | [[Image:Memb localization.jpg|300px]] | ||
Revision as of 15:41, 24 January 2015
| |||||||||||