User:Braden Sciarra/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 16: | Line 16: | ||
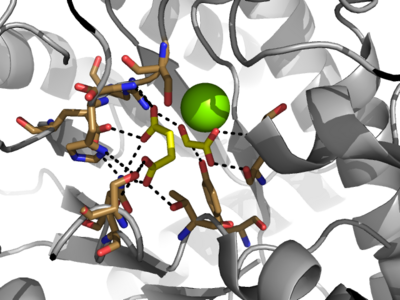
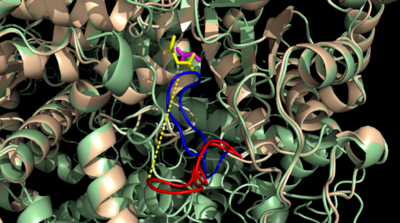
The <scene name='69/697526/Catalytic_loop/1'>Catalytic Loop</scene> of the Isocitrate Lyase enzyme is composed of residues 185-196, and can exist in both the open and closed conformation. In the open conformation, the catalytic loop is oriented such that the catalytic CYS191 residue is located far from the active site, allowing for solvent accessibility and substrate binding. Upon substrate binding, the catalytic loop adopts a closed conformation, moving between ten and fifteen angstroms<ref name="ICL">PMID:10932251</ref>. This closed conformation will cause the binding site to become inaccessible to the solvent. The loop closure is triggered by the movement of the Mg2+ ion that occurs upon binding of the succinate. This movement of the Mg2+ ion results in electrostatic interactions at LYS189, causing the loop to close. | The <scene name='69/697526/Catalytic_loop/1'>Catalytic Loop</scene> of the Isocitrate Lyase enzyme is composed of residues 185-196, and can exist in both the open and closed conformation. In the open conformation, the catalytic loop is oriented such that the catalytic CYS191 residue is located far from the active site, allowing for solvent accessibility and substrate binding. Upon substrate binding, the catalytic loop adopts a closed conformation, moving between ten and fifteen angstroms<ref name="ICL">PMID:10932251</ref>. This closed conformation will cause the binding site to become inaccessible to the solvent. The loop closure is triggered by the movement of the Mg2+ ion that occurs upon binding of the succinate. This movement of the Mg2+ ion results in electrostatic interactions at LYS189, causing the loop to close. | ||
| - | == Mechanism == | + | ==Mechanism== |
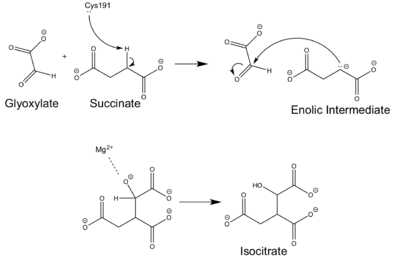
[[Image:ICL Mechanism.png|400 px|right|thumb|Chemical Mechanism of Isocitrate Lyase]] | [[Image:ICL Mechanism.png|400 px|right|thumb|Chemical Mechanism of Isocitrate Lyase]] | ||
Isocitrate lyase catalyzes a reversible aldol condensation, converting isocitrate to glyoxylate and succinate via the breaking of a C-C bond. Within the active site of ICL the HIS193 residue deprotonates the CYS191 residue of the active site in order to increase its basicity<ref name="ICL">PMID:10932251</ref>. The Cys 191 residue then deprotonates the alpha carbon adjacent to one of the carbonyl groups of succinate, thus forming the enolic intermediate. The negatively charged alpha carbon atom of the enolic intermediate acts as a nucleophile that attacks the carbonyl carbon of the aldehyde of glyoxylate. The nucleophilic attack will place a negative charge on the oxygen atom oxygen at the former carbonyl oxygen of the aldehyde, which will be stabilized by positive charges of the Mg2+ ion, Arg 228 and His 180<ref name="ICL">PMID:10932251</ref>. The protonation of this species will yield the final product. It is important to note that this reaction is entirely reversible; the breakdown of isocitrate into glyoxylate and succinate occurs using a similar mechanism. | Isocitrate lyase catalyzes a reversible aldol condensation, converting isocitrate to glyoxylate and succinate via the breaking of a C-C bond. Within the active site of ICL the HIS193 residue deprotonates the CYS191 residue of the active site in order to increase its basicity<ref name="ICL">PMID:10932251</ref>. The Cys 191 residue then deprotonates the alpha carbon adjacent to one of the carbonyl groups of succinate, thus forming the enolic intermediate. The negatively charged alpha carbon atom of the enolic intermediate acts as a nucleophile that attacks the carbonyl carbon of the aldehyde of glyoxylate. The nucleophilic attack will place a negative charge on the oxygen atom oxygen at the former carbonyl oxygen of the aldehyde, which will be stabilized by positive charges of the Mg2+ ion, Arg 228 and His 180<ref name="ICL">PMID:10932251</ref>. The protonation of this species will yield the final product. It is important to note that this reaction is entirely reversible; the breakdown of isocitrate into glyoxylate and succinate occurs using a similar mechanism. | ||
| + | |||
| + | ==Elucidation of ICL Structure Using Inhibitors== | ||
| + | |||
| + | The two inhibitors used to elucidate the structure of ICL were 3-nitropropionate and 3-bromopyruvate. The 3-nitropropionate was used to mimic the succinate, while the 3-bromopyruvate is used to mimic the glyoxylate. These two inhibitors have also been shown to be good inhibitors of isocitrate lyase in ''M. avium'' indicating that their inhibitory capacity is conserved across multiple species. A mutant isocitrate lyase C191S, in conjunction with the aforementioned substrate mimics, was used to elucidate the first high resolution crystal structure of ICL. Dehalogenated 3-nitropropionate works to inhibit isocitrate lyase by forming a covalent bond with the Ser 191 residue in the active site. This 3-nitropropionate occupies the same site that the succinate would occupy. The C191S mutant adopts a conformation almost identical to the Cys 191 residue in the wild type indicating that this is an accurate depiction of the conformation. | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 23:55, 7 April 2015
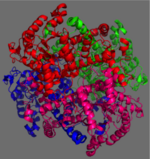
Isocitrate Lyase from Mycobacterium Tuberculosis
| |||||||||||