This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1074
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
== '''General Structural Information''' == | == '''General Structural Information''' == | ||
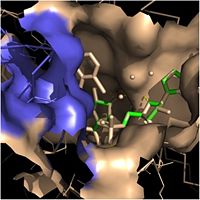
| - | Crystal structures of InhA reveal a <scene name='69/694241/Homotetramer_subunits_labeled/1'>homotetramer</scene> (each subunit featured with a different color) in aqueous solution with separate ligand binding sites in each subunit. Each <scene name='69/694241/Monomer_subunit_no_ligands/1'>monomer</scene> subunit is composed of 289 residues and features a typical [http://en.wikipedia.org/wiki/Rossmann_fold Rossmann fold] containing a single NADH binding site. The <scene name='69/694241/Secondary_structure_black/1'>secondary structure</scene> of InhA is made up of several alpha helices (pink), beta sheets (gold), and beta turns (white). This enzyme also features a fatty acyl binding crevice that accommodates the long-chain fatty acyl substrate needed to synthesize mycolic acid precursors. The <scene name='69/694241/Helix6_helix7_updated/1'>alpha-6 and alpha-7 helices</scene> of the InhA form one side of the fatty acyl binding crevice, referred to as the <scene name='69/694241/Monomer_subunit_196_219/1'> substrate binding loop</scene> (residues 196-219). | + | Crystal structures of InhA reveal a <scene name='69/694241/Homotetramer_subunits_labeled/1'>homotetramer</scene> (each subunit featured with a different color) in aqueous solution with separate ligand binding sites in each subunit. Each <scene name='69/694241/Monomer_subunit_no_ligands/1'>monomer</scene> subunit is composed of 289 residues and features a typical [http://en.wikipedia.org/wiki/Rossmann_fold Rossmann fold] containing a single NADH binding site. The <scene name='69/694241/Secondary_structure_black/1'>secondary structure</scene> of InhA is made up of several alpha helices (pink), beta sheets (gold), and beta turns (white). This enzyme also features a fatty acyl binding crevice that accommodates the long-chain fatty acyl substrate needed to synthesize mycolic acid precursors. [[Image:Fatty Acyl Binding Crevice.jpg|thumb|200px|left|Fatty Acyl Binding Crevice (substrate binding loop in purple; substrates pictured inside the crevice)]] The <scene name='69/694241/Helix6_helix7_updated/1'>alpha-6 and alpha-7 helices</scene> of the InhA form one side of the fatty acyl binding crevice, referred to as the <scene name='69/694241/Monomer_subunit_196_219/1'> substrate binding loop</scene> (residues 196-219). One side of this crevice is open and exposed to solvent, which allows the substrates to access the binding pocket of this enzyme. The size of the substrate binding loop is a primary determinant of the ability of InhA to select for fatty acyl chains longer than 16 carbons to successfully produce mycolic acid precursors. |
| - | + | ||
==='''Ligands'''=== | ==='''Ligands'''=== | ||
| Line 25: | Line 24: | ||
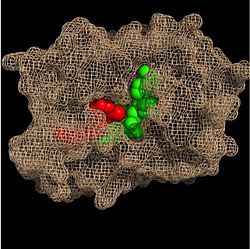
Within the fatty acyl binding crevice, the NADH substrate sits on the top shelf of the Rossmann fold, and the fatty acyl substrate sits on top of the NADH substrate. Due to its position within the crevice, the ''trans'' double bond of the fatty acyl substrate is found near the closed end of the crevice and is located directly adjacent to the nicotinamide ring of [http://en.wikipedia.org/wiki/Nicotinamide_adenine_dinucleotide NADH]. [[Image:Binding Pocket - Mesh.jpg|thumb|250px|right|Substrate Binding Pocket (NADH in green; fatty acyl substrate in red)]] These two molecules interact with each other via hydrogen bonding. The first hydrogen bond occurs between the phosphate oxygen of NADH and the amide nitrogen of the N-acetylcysteamine portion of the fatty acyl substrate. The second hydrogen bond occurs between the 2'-hydroxyl of the nicotinamide ribose of NADH and the thioester carbonyl oxygen of the fatty acyl substrate. | Within the fatty acyl binding crevice, the NADH substrate sits on the top shelf of the Rossmann fold, and the fatty acyl substrate sits on top of the NADH substrate. Due to its position within the crevice, the ''trans'' double bond of the fatty acyl substrate is found near the closed end of the crevice and is located directly adjacent to the nicotinamide ring of [http://en.wikipedia.org/wiki/Nicotinamide_adenine_dinucleotide NADH]. [[Image:Binding Pocket - Mesh.jpg|thumb|250px|right|Substrate Binding Pocket (NADH in green; fatty acyl substrate in red)]] These two molecules interact with each other via hydrogen bonding. The first hydrogen bond occurs between the phosphate oxygen of NADH and the amide nitrogen of the N-acetylcysteamine portion of the fatty acyl substrate. The second hydrogen bond occurs between the 2'-hydroxyl of the nicotinamide ribose of NADH and the thioester carbonyl oxygen of the fatty acyl substrate. | ||
| - | |||
==='''Substrate Binding Loop Flexibility''' === | ==='''Substrate Binding Loop Flexibility''' === | ||
In addition to the hydrogen bonding that occurs between the NADH and fatty acyl substrate within the crevice, each of these molecules is also held in place within the crevice through interactions with the side chains of surrounding <scene name='69/694241/Sbl_hydrophobic/1'>hydrophobic</scene> (purple) residues. The majority of these <scene name='69/694241/Hydrophobic_residues/1'>hydrophobic</scene> residues anchoring the substrates are found within the substrate binding loop itself, including Ala-198, Met-199, Ala-201, Ile-202, Leu-207, Ile-215, and Leu-218. Additional hydrophobic amino acids that are not a part of the substrate binding loop also play a role in positioning and stabilizing the fatty acyl chain in the crevice. Studies have found that the fatty acyl substrate adopts a u-shaped conformation to facilitate binding. As the fatty acyl substrate binds in the crevice, the substrate binding loop shifts outward toward the solvent, and the [[Tyr-158]] residue is rotated to facilitate the binding of fatty acyl chains of 16 carbons or greater. During this conformational change upon substrate binding, no hydrogen bonds are broken, which supports the flexibility of the substrate binding loop. It is likely that this flexibility of the substrate binding loop provides increased freedom for the active site to accept fatty acyl substrates of varying carbon chain lengths. | In addition to the hydrogen bonding that occurs between the NADH and fatty acyl substrate within the crevice, each of these molecules is also held in place within the crevice through interactions with the side chains of surrounding <scene name='69/694241/Sbl_hydrophobic/1'>hydrophobic</scene> (purple) residues. The majority of these <scene name='69/694241/Hydrophobic_residues/1'>hydrophobic</scene> residues anchoring the substrates are found within the substrate binding loop itself, including Ala-198, Met-199, Ala-201, Ile-202, Leu-207, Ile-215, and Leu-218. Additional hydrophobic amino acids that are not a part of the substrate binding loop also play a role in positioning and stabilizing the fatty acyl chain in the crevice. Studies have found that the fatty acyl substrate adopts a u-shaped conformation to facilitate binding. As the fatty acyl substrate binds in the crevice, the substrate binding loop shifts outward toward the solvent, and the [[Tyr-158]] residue is rotated to facilitate the binding of fatty acyl chains of 16 carbons or greater. During this conformational change upon substrate binding, no hydrogen bonds are broken, which supports the flexibility of the substrate binding loop. It is likely that this flexibility of the substrate binding loop provides increased freedom for the active site to accept fatty acyl substrates of varying carbon chain lengths. | ||
| - | |||
==='''Importance of Tyr-158'''=== | ==='''Importance of Tyr-158'''=== | ||
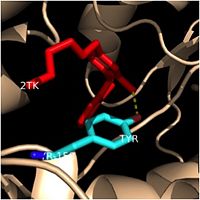
| - | One example of a hydrophobic amino acid that is not a part of the substrate binding loop yet interacts with the fatty acyl substrate is [[Tyr-158]]. [[Image:Tyr-158.jpg|thumb|200px|left|Tyr-158 ( | + | One example of a hydrophobic amino acid that is not a part of the substrate binding loop yet interacts with the fatty acyl substrate is [[Tyr-158]]. [[Image:Tyr-158.jpg|thumb|200px|left|Tyr-158 (light blue) hydrogen bonding to the fatty acyl substrate, 2TK (red)]] This amino acid is conserved in other enoyl-ACP reductases in both bacteria and plants, so it likely plays an essential role in the function of these specific enzymes. Studies have shown that Tyr-158 forms the only direct hydrogen bond that exists between the InhA protein and the fatty acyl substrate. This hydrogen bond occurs between the hydroxyl oxygen on the side chain of Tyr-158 and the thioester carbonyl oxygen of the fatty acyl substrate. As a result of the highly conserved nature and the specific hydrogen bonding capabilities of this amino acid, it is likely that Tyr-158 plays an important role in fatty acyl substrate binding in enoyl-ACP reductases. |
Revision as of 02:52, 8 April 2015
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
Enoyl-ACP Reductase InhA from Mycobacterium tuberculosis
| |||||||||||