This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1058
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
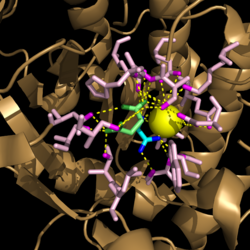
[[Image:Active_Site_Hydrogen_Bonding.png|250 px|left|thumb|'''Figure 3. Active site residues hydrogen bound to a cofactor and the products of the catalyzed isocitrate reaction.''' Glyoxylate is shown in blue, succinate is shown in green, and the Mg<sup>2+</sup> cofactor is shown in yellow.]] | [[Image:Active_Site_Hydrogen_Bonding.png|250 px|left|thumb|'''Figure 3. Active site residues hydrogen bound to a cofactor and the products of the catalyzed isocitrate reaction.''' Glyoxylate is shown in blue, succinate is shown in green, and the Mg<sup>2+</sup> cofactor is shown in yellow.]] | ||
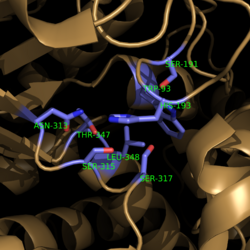
| - | [[Image:Active Site Residues.png|250 px| | + | [[Image:Active Site Residues.png|250 px|center|thumb|'''Figure 4. Active Site Residues.''' All eight active site residues necessary for catalysis of isocitrate are shown in slate. However, the protein shown is a C191S mutant of isocitrate lyase.]] |
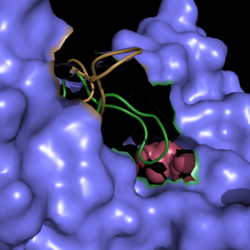
[[Image:Active Loop Shift.png|250 px|left|thumb|'''Figure 2. Active Site Loop Shift.''' Binding of the ligand to the enzyme results in a conformational shift that facilitates the breakdown of isocitrate. The active site loop unbound is shown in wheat and the active site loop bound is shown in green.]] | [[Image:Active Loop Shift.png|250 px|left|thumb|'''Figure 2. Active Site Loop Shift.''' Binding of the ligand to the enzyme results in a conformational shift that facilitates the breakdown of isocitrate. The active site loop unbound is shown in wheat and the active site loop bound is shown in green.]] | ||
| Line 25: | Line 25: | ||
==Mechanism of Action== | ==Mechanism of Action== | ||
| - | [[Image:TCA_Cycle.png|500 px| | + | [[Image:TCA_Cycle.png|500 px|left|thumb|'''Figure 5. Citric Acid Cycle with Glyoxylate Shunt Pathway.''' In several bacterial species, there is a carbon conserving gloxylate shunt pathway that converts isocitrate to malate in two steps instead of the usual five steps.]] |
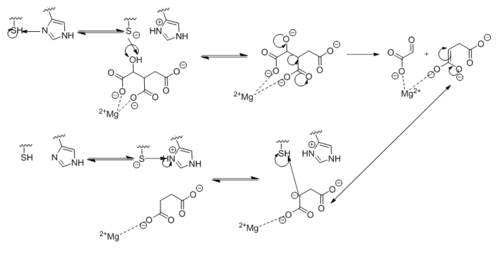
| - | [[Image:Complete_Mechanism.PNG|500 px| | + | [[Image:Complete_Mechanism.PNG|500 px|left|thumb|'''Figure 6. Observed Mechanism for the Breakdown of Isocitrate by Isocitrate Lyase.''' His193 shifts the pKa of Cys191 and removes its proton. This allows Cys191 to extract a proton from the hydroxyl group of isocitrate. The resulting oxyanion forms a carbonyl and forces the lysis of a C-C bond. Glyoxylate and the enol form of succinate are formed and stabilized with a Mg<sup>2+</sup> ion. The succinate enolate resonates and extracts the proton back from Cys191 to form succinate.]] |
| + | |||
Revision as of 03:01, 10 April 2015
Isocitrate Lyase from Mycobacterium tuberculosis
| |||||||||||
References
- ↑ Cozzone, A.; Regulation of acetate metabolism by protein phosphorylation in enteric bacteria. Annual Review of Microbiology. 1998, 52:127-164. doi: 10.1146/annurev.micro.52.1.127.
- ↑ Srivastava, V.; Janin, A.; Srivastava, B.; Srivastava, R.; Selection of genes of Mycobacterium tuberculosis upregulated during residence in lungs of infected mice. ScienceDirect. 2007. doi:10.1016/j.tube.2007.10.002.
- ↑ Muñoz-Elías, E.; McKinney, J.; M. tuberculosis isocitrate lyases 1 and 2 are jointly required for in vivo growth and virulence. Nat. Med. 2005. 11(6):638-644. doi:10.1038/nm1252.
- ↑ Dunn, M.; Ramírez-Trujillo, J.; Hernández-Lucas, I.; Major roles of isocitrate lyase and malate synthase in bacterial and fungal pathogenesis. Microbiology. 2009. 155:3166-3175. doi:10.1099/mic.0.030858-0.