Sandbox Reserved 1061
From Proteopedia
(Difference between revisions)
| Line 35: | Line 35: | ||
== Relevance == | == Relevance == | ||
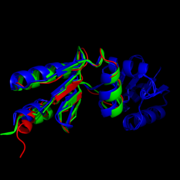
| - | Like most NrdHs, MtNrdH is similar in sequence to glutaredoxins, but structurally similar to thioredoxins. MtNrdH also accepts electrons from | + | Like most NrdHs, MtNrdH is similar in sequence to glutaredoxins, but structurally similar to thioredoxins. MtNrdH also accepts electrons from thioredoxin reductase, a characteristic of thioredoxins, but not glutaredoxins.<ref>Swastik, Phulera and Mande, Shekhar C. (2013) 4057.</ref> [[Image:Image-Super imposed molecules.png|thumb|left|Structural comparison of NrdHs with "thioredoxin folds": ''E. Coli'' NrdH (green), ''C. ammoniagenes'' NrdH (blue), ''M. tuberculosis'' NrdH (red)]]<ref>DOI 10.1002/ijch.201300024</ref> <ref>PMID:21638687</ref> |
| - | Similar structures of NrdH have been isolated in other primitive species including ''E. coli'', ''S. pyogenes'', ''S. typhimurium'', ''D. deserti'', ''S. flexneri 2457T'', and ''S. dysenteriae''. In higher order multi-cellular organisms, however the NrdH protein is replaced by more complex glutaredoxins or | + | Similar structures of NrdH have been isolated in other primitive species including ''E. coli'', ''S. pyogenes'', ''S. typhimurium'', ''D. deserti'', ''S. flexneri 2457T'', and ''S. dysenteriae''. In higher order multi-cellular organisms, however the NrdH protein is replaced by more complex glutaredoxins or thioredoxins. This observation leads some to speculate that NrdH is one of the very first ancestors in the ribonucleotide reduction pathway. <ref>Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.</ref> If this is true, NrdH can be seen as a critical protein that allowed for the development of DNA-based life since deoxyribonucleotides could not have existed without the ribonucleotide reduction pathway. A better understanding of the evolutionary timeline of NrdH and similar proteins could shed greater light onto the RNA Wold Hypothesis, specifically describing the time frame of emergence of DNA based life. |
Revision as of 19:09, 10 April 2015
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
Structure of Mycobacterium Tuberculosis NrdH
| |||||||||||
References
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.
- ↑ "Tuberculosis." Media Centre. World Health Organization, Web. 16 Mar. 2015. Media Centre. <http://www.who.int/mediacentre/factsheets/fs104/en/>.
- ↑ "Tuberculosis." Media Centre. World Health Organization, Web. 16 Mar. 2015. Media Centre. <http://www.who.int/mediacentre/factsheets/fs104/en/>.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) 4060.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) 4057.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) 4056.
- ↑ Nelson, David L., and Michael M. Cox. Lehninger Principles of Biochemistry. 5th ed. New York: W.H. Freeman, 2008. 888-889.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) 4056.
- ↑ Makhlynets, O., Boal, A. K., Rhodes, D. V., Kitten, T., Rosenzweig, A. C., & Stubbe, J. (2014). Streptococcus sanguinis Class Ib Ribonucleotide Reductase: HIGH ACTIVITY WITH BOTH IRON AND MANGANESE COFACTORS AND STRUCTURAL INSIGHTS. The Journal of Biological Chemistry, 289(9), 6259–6272. doi:10.1074/jbc.M113.533554.
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) 4057.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Swastik, Phulera and Mande, Shekhar C. (2013) The Crystal Structure of Mycobacterium tuberculosis NrdH at 0.87Å Suggests a Possible Mode of Its Activity. Biochemistry 52, 4056-4065.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644