This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1066
From Proteopedia
| Line 27: | Line 27: | ||
=Structure = | =Structure = | ||
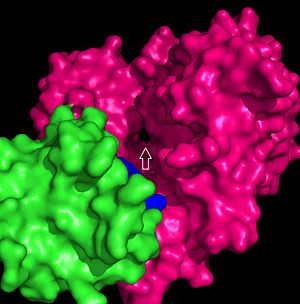
FadD13 is composed of 503 amino acid residues divided into three main regions: The <scene name='69/694233/N_terminal_domain/1'>N-terminal domain</scene> (residues 1-395) and <scene name='69/694233/C-terminal_domain/2'>C-terminal domain</scene> (residues 402-503) which are connected via a flexible <scene name='69/694233/Linker_section/2'>linker</scene> represented in dark blue (residues 396-401).<ref name="Our Paper"/> Each region plays an important role in the activation of fatty acids. The large N-terminal domain houses many key structural features involved in fatty acid activation, but ultimately it is the flexible linker that allows movement of the C-terminal domain to from the fully functioning active site of FadD13 (Figure 1). | FadD13 is composed of 503 amino acid residues divided into three main regions: The <scene name='69/694233/N_terminal_domain/1'>N-terminal domain</scene> (residues 1-395) and <scene name='69/694233/C-terminal_domain/2'>C-terminal domain</scene> (residues 402-503) which are connected via a flexible <scene name='69/694233/Linker_section/2'>linker</scene> represented in dark blue (residues 396-401).<ref name="Our Paper"/> Each region plays an important role in the activation of fatty acids. The large N-terminal domain houses many key structural features involved in fatty acid activation, but ultimately it is the flexible linker that allows movement of the C-terminal domain to from the fully functioning active site of FadD13 (Figure 1). | ||
| - | |||
| - | |||
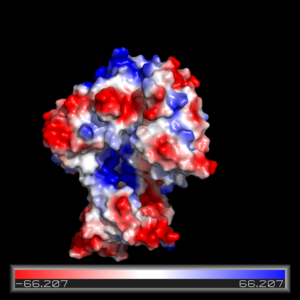
== Electrostatics == | == Electrostatics == | ||
| Line 38: | Line 36: | ||
== Hydrophobic Tunnel == | == Hydrophobic Tunnel == | ||
| - | The <scene name='69/694233/Hydrophobic_tunnel/ | + | The <scene name='69/694233/Hydrophobic_tunnel/2'>hydrophobic tunnel</scene> of FadD13 is essential to the transport and accommodation of very long fatty acids from the membrane into the cell. This tunnel runs through the middle of FadD13 from the arginine rich lid loop to the ATP binding site and is situated between the and alpha helices α8-α9 and parallel beta sheet β9- β14 (Figure 4).<ref name="Our Paper"/> Negatively charged residues at the active site of FadD13 are the driving factor in the attraction of the fatty acid from the membrane through the hydrophobic tunnel of the enzyme. |
[[Image:Hydrophobic tunnel 2.jpg|300 px|left|thumb|Figure 4: Pmyol depiction of hydrophobic tunnel.]] | [[Image:Hydrophobic tunnel 2.jpg|300 px|left|thumb|Figure 4: Pmyol depiction of hydrophobic tunnel.]] | ||
| Line 63: | Line 61: | ||
==References== | ==References== | ||
<references/> | <references/> | ||
| - | |||
Revision as of 17:46, 17 April 2015
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
Mycobacterium tuberculosis very-long-chain fatty acyl-CoA synthetase
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 Andersson CS, Lundgren CA, Magnusdottir A, Ge C, Wieslander A, Molina DM, Hogbom M. The Mycobacterium tuberculosis Very-Long-Chain Fatty Acyl-CoA Synthetase: Structural Basis for Housing Lipid Substrates Longer than the Enzyme. Structure. 2012 May 2. PMID:22560731 doi:10.1016/j.str.2012.03.012
- ↑ Jatana N, Jangid S, Khare G, Tyagi AK, Latha N. Molecular modeling studies of Fatty acyl-CoA synthetase (FadD13) from Mycobacterium tuberculosis--a potential target for the development of antitubercular drugs. J Mol Model. 2011 Feb;17(2):301-13. doi: 10.1007/s00894-010-0727-3. Epub 2010 May, 8. PMID:20454815 doi:http://dx.doi.org/10.1007/s00894-010-0727-3
- ↑ 3.0 3.1 3.2 3.3 3.4 Khare G, Gupta V, Gupta RK, Gupta R, Bhat R, Tyagi AK. Dissecting the role of critical residues and substrate preference of a Fatty Acyl-CoA Synthetase (FadD13) of Mycobacterium tuberculosis. PLoS One. 2009 Dec 21;4(12):e8387. doi: 10.1371/journal.pone.0008387. PMID:20027301 doi:10.1371/journal.pone.0008387
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 Jatana N, Jangid S, Khare G, Tyagi AK, Latha N. Molecular modeling studies of Fatty acyl-CoA synthetase (FadD13) from Mycobacterium tuberculosis--a potential target for the development of antitubercular drugs. J Mol Model. 2011 Feb;17(2):301-13. doi: 10.1007/s00894-010-0727-3. Epub 2010 May, 8. PMID:20454815 doi:http://dx.doi.org/10.1007/s00894-010-0727-3
- ↑ 5.0 5.1 5.2 Schroeder EK, de Souza N, Santos DS, Blanchard JS, Basso LA. Drugs that inhibit mycolic acid biosynthesis in Mycobacterium tuberculosis. Curr Pharm Biotechnol. 2002 Sep;3(3):197-225. PMID:12164478
External Resources
Lipids Wikipedia page
Tuberculosis Wikipedia page
Mycobacterium tuberculosis Wikipedia page
Coenzyme A Wikipedia page
Acyl CoA Wikipedia Page
Mycolic Acid Wikipedia page
Peripheral Membrane Protein Wikipedia page
Ethionamide Wikipedia page
Isoniazid Wikipedia page
Thiocarlide Wikipedia page
Pyrazinamide Wikipedia page