This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1070
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
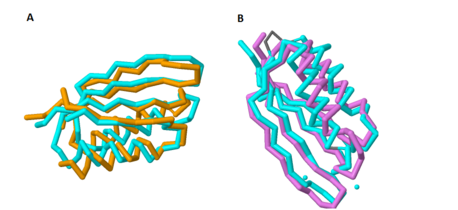
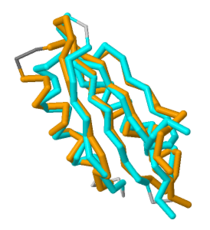
This domain of MgtC, in contrast, is highly variable in comparison to several orthologs, as presented by Yang ''et al''. However, it is this <scene name='69/698113/Secondary_structured_coloring/2'>tertiary structure</scene> containing two α-helices and four anti-parallel β-sheets that is incredibly indicative of the MgtC super family. Through a sequence alignment of five known functional MgtC orthologs from [http://en.wikipedia.org/wiki/Pathogen pathogens] that survive inside [http://en.wikipedia.org/wiki/Macrophage macrophages] (''M. tuberculosis, [http://en.wikipedia.org/wiki/Brucella_melitensis B. melitensis], [http://en.wikipedia.org/wiki/Burkholderia_cenocepacia B. cenocepacia], [http://en.wikipedia.org/wiki/Yersinia_pestis Y. pestis],'' and ''[http://en.wikipedia.org/wiki/Salmonella_enterica_subsp._enterica S. Typhimurium]''), seven strictly conserved residues were found to be scattered along the whole sequence of the relatively hydrophilic and soluble C-terminal domain. <ref name="mgtc"/> | This domain of MgtC, in contrast, is highly variable in comparison to several orthologs, as presented by Yang ''et al''. However, it is this <scene name='69/698113/Secondary_structured_coloring/2'>tertiary structure</scene> containing two α-helices and four anti-parallel β-sheets that is incredibly indicative of the MgtC super family. Through a sequence alignment of five known functional MgtC orthologs from [http://en.wikipedia.org/wiki/Pathogen pathogens] that survive inside [http://en.wikipedia.org/wiki/Macrophage macrophages] (''M. tuberculosis, [http://en.wikipedia.org/wiki/Brucella_melitensis B. melitensis], [http://en.wikipedia.org/wiki/Burkholderia_cenocepacia B. cenocepacia], [http://en.wikipedia.org/wiki/Yersinia_pestis Y. pestis],'' and ''[http://en.wikipedia.org/wiki/Salmonella_enterica_subsp._enterica S. Typhimurium]''), seven strictly conserved residues were found to be scattered along the whole sequence of the relatively hydrophilic and soluble C-terminal domain. <ref name="mgtc"/> | ||
| - | A large hydrophobic core has conserved residues <scene name='69/698113/ | + | A large hydrophobic core has conserved residues <scene name='69/698113/Colored_core_residues/6'>Cysteine-155, Arginine-164, Glutamine-160, and Alanine-195</scene>. |
[[Image:Final_Final_Core_Logo.PNG |625× 116px|thumb|left|Four strictly conserved residues of five known functional MgtC orthologs of the soluble C-terminal domain. | [[Image:Final_Final_Core_Logo.PNG |625× 116px|thumb|left|Four strictly conserved residues of five known functional MgtC orthologs of the soluble C-terminal domain. | ||
| Line 33: | Line 33: | ||
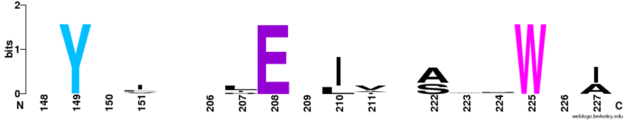
| - | The opposite side of the protein has a small cluster of conserved residues <scene name='69/698113/Conserved_surface_residues/ | + | The opposite side of the protein has a small cluster of conserved residues <scene name='69/698113/Conserved_surface_residues/6'>Tyrosine-149, Glutamine-208, and Tryptophan-225</scene>. |
[[Image:Final_Surface_Web_Logo.PNG |625× 121px|thumb|left|Four strictly conserved residues of five known functional MgtC orthologs of the soluble C-terminal domain.The figure was prepared using WebLogo. (http://weblogo.berkeley.edu/)]] | [[Image:Final_Surface_Web_Logo.PNG |625× 121px|thumb|left|Four strictly conserved residues of five known functional MgtC orthologs of the soluble C-terminal domain.The figure was prepared using WebLogo. (http://weblogo.berkeley.edu/)]] | ||
Revision as of 21:35, 17 April 2015
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
MgtC: A Virulence Factor From Mycobacterium tuberculosis
| |||||||||||
References
- ↑ Singh, G.; Singh, G.; Jadeja, D.; Kaur, J. Lipid hydrolyzing enzymes in virulence: Mycobacterium tuberculosis as a model system. Critical Reviews in Microbiology 2010, 36(3): 259-269. DOI: 10.3109/1040841X.2010.482923.
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 Yang, Y.; Labesse, G.; Carrere-Kremer, S.; Esteves, K.; Kremer, L.; Cohen-Gonsaud, M.; Blanc-Potard, A. The C-terminal domain of the virulence factor mgtc is a divergent act domain. J Bacteriol. 2012, 194(22): 6255-6263. DOI: 10.1128/JB.01424-12.
- ↑ Blanc-Potard, A.B.; Lafay, B. MgtC as a horizontally-acquired virulence factor of intracellular bacterial pathogens : evidence from molecular phylogeny and comparative genomics. J Mol Evol. 2003, 57(4): 479-86. DOI: 10.1007/s00239-003-2496-4
- ↑ Belon, C.; Gannoun-Zaki, L.; Lutfalla, G.; Kremer, L.; Blanc-Potard, A.B. Mycobacterium marinum mgtc plays a role in phagocytosis but is dispensable for intracellular multiplication. Plos One 2014, 1-23. DOI: 10.1371/journal.pone.0116052.
- ↑ 5.0 5.1 5.2 Jean-Francois, F.L.; Dai, J.; Yu, L. ; Myrick, A. ; Rubin, E. ; et al. Binding of mgtr, a salmonella transmembrane regulatory peptide, to mgtc, a mycobacterium tuberculosis virulence factor: a structural study. DOI:10.1016/j.jmb.2013.10.014