This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1063
From Proteopedia
(Difference between revisions)
| Line 21: | Line 21: | ||
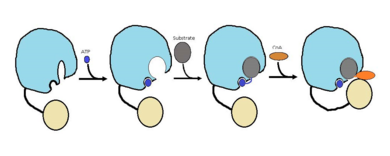
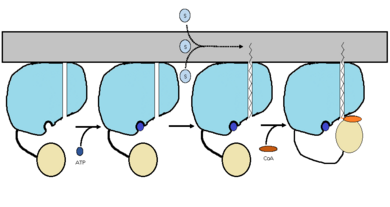
The FadD13 enzyme functions to activate lipids. Once the lipids are activated, they can continue on into metabolic pathways. This is done by ATP/AMP binding to the <scene name='69/694230/Fadd13_subunits/12'>ATP/AMP binding region</scene>. Once ATP/AMP is bound, the long lipid chain up to 26 carbons may bind in the <scene name='69/694230/Fadd13_subunits/13'>hydrophobic tunnel</scene> of the enzyme. Upon binding of the substrate, the C terminal swings up to close off the tunnel. From there CoA can bind to produce the final product, an acyl-CoA Thioester. The lipid can now move transversely throughout the membrane and throughout the rest of the cell. Below is the proposed mechanism for ACSVL proteins. | The FadD13 enzyme functions to activate lipids. Once the lipids are activated, they can continue on into metabolic pathways. This is done by ATP/AMP binding to the <scene name='69/694230/Fadd13_subunits/12'>ATP/AMP binding region</scene>. Once ATP/AMP is bound, the long lipid chain up to 26 carbons may bind in the <scene name='69/694230/Fadd13_subunits/13'>hydrophobic tunnel</scene> of the enzyme. Upon binding of the substrate, the C terminal swings up to close off the tunnel. From there CoA can bind to produce the final product, an acyl-CoA Thioester. The lipid can now move transversely throughout the membrane and throughout the rest of the cell. Below is the proposed mechanism for ACSVL proteins. | ||
| + | |||
| + | ==Future Work== | ||
==Relevant Pages== | ==Relevant Pages== | ||
| - | [http://proteopedia.org/wiki/index.php/3t5b 3T5B] | + | * [http://proteopedia.org/wiki/index.php/3t5b 3T5B] |
| - | [http://proteopedia.org/wiki/index.php/3t5c 3T5C] | + | * [http://proteopedia.org/wiki/index.php/3t5c 3T5C] |
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
{{reflist}} | {{reflist}} | ||
Revision as of 18:12, 21 April 2015
FadD13
| |||||||||||
References
- ↑ Watkins PA, Maiguel D, Jia Z, Pevsner J. Evidence for 26 distinct acyl-coenzyme A synthetase genes in the human genome. J Lipid Res. 2007 Dec;48(12):2736-50. Epub 2007 Aug 30. PMID:17762044 doi:http://dx.doi.org/M700378-JLR200
- ↑ Kochan G, Pilka ES, von Delft F, Oppermann U, Yue WW. Structural snapshots for the conformation-dependent catalysis by human medium-chain acyl-coenzyme A synthetase ACSM2A. J Mol Biol. 2009 May 22;388(5):997-1008. Epub 2009 Apr 1. PMID:19345228 doi:10.1016/j.jmb.2009.03.064
- ↑ 3.0 3.1 3.2 3.3 3.4 Andersson CS, Lundgren CA, Magnusdottir A, Ge C, Wieslander A, Molina DM, Hogbom M. The Mycobacterium tuberculosis Very-Long-Chain Fatty Acyl-CoA Synthetase: Structural Basis for Housing Lipid Substrates Longer than the Enzyme. Structure. 2012 May 2. PMID:22560731 doi:10.1016/j.str.2012.03.012
- ↑ 4.0 4.1 Khare G, Gupta V, Gupta RK, Gupta R, Bhat R, Tyagi AK. Dissecting the role of critical residues and substrate preference of a Fatty Acyl-CoA Synthetase (FadD13) of Mycobacterium tuberculosis. PLoS One. 2009 Dec 21;4(12):e8387. doi: 10.1371/journal.pone.0008387. PMID:20027301 doi:10.1371/journal.pone.0008387