This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Heat Shock Proteins
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | + | <StructureSection load='' size='450' side='right' scene='Journal:JBSD:30/Cv/5' caption=''> | |
[[Heat Shock Proteins]] (Hsp). The Hsp are proteins which are expressed when cells are exposed to stresses like heat. Hsp60, Hsp70, Hsp90 etc. are named according to their molecular weights. Their function is to provide protection from stress. Hsp contains a nucleotide-binding domain (NBD), a substrate-binding domain (sub) and C-terminal lid domain (lid). In E. coli there are additional Hsp – HslV (19kD), HslJ and HslU (50kD). The heat-inducible Hsp is called '''heat shock cognate protein''' (Hsc). Hsp70 from E. coli is called '''DnaK'''. | [[Heat Shock Proteins]] (Hsp). The Hsp are proteins which are expressed when cells are exposed to stresses like heat. Hsp60, Hsp70, Hsp90 etc. are named according to their molecular weights. Their function is to provide protection from stress. Hsp contains a nucleotide-binding domain (NBD), a substrate-binding domain (sub) and C-terminal lid domain (lid). In E. coli there are additional Hsp – HslV (19kD), HslJ and HslU (50kD). The heat-inducible Hsp is called '''heat shock cognate protein''' (Hsc). Hsp70 from E. coli is called '''DnaK'''. | ||
| Line 9: | Line 9: | ||
* [[Molecular Playground/sHSP]] for small Hsp | * [[Molecular Playground/sHSP]] for small Hsp | ||
| - | === Conformational dynamics of full-length inducible human Hsp70 derived from microsecond molecular dynamics simulations in explicit solvent | + | === Conformational dynamics of full-length inducible human Hsp70 derived from microsecond molecular dynamics simulations in explicit solvent <ref>doi 10.1080/07391102.2012.726190</ref> === |
| - | + | ||
| - | + | ||
| - | + | ||
'''1. Hsp70, molecular chaperone''' | '''1. Hsp70, molecular chaperone''' | ||
| Line 38: | Line 35: | ||
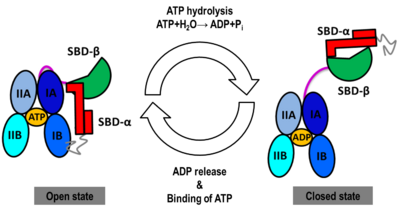
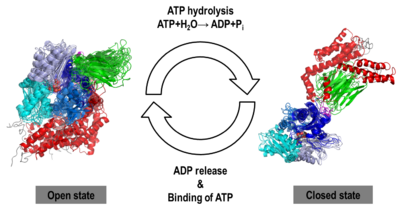
We found that human Hsp70, <scene name='Journal:JBSD:30/Cv/13'>both in its open and closed states</scene>, was better represented by an ensemble of conformations on a μs timescale, corresponding to different local minima of its free-energy landscape in good agreement with low-resolution experimental data of Hsp70s homologs (Small Angle Xray-Scattering and single-molecule Förster Resonance Energy Transfer) to which the MD simulations were compared in details. These structural ensembles represent the first attempt to model the conformational heterogeneity of a full-length molecular chaperone at the atomic scale. | We found that human Hsp70, <scene name='Journal:JBSD:30/Cv/13'>both in its open and closed states</scene>, was better represented by an ensemble of conformations on a μs timescale, corresponding to different local minima of its free-energy landscape in good agreement with low-resolution experimental data of Hsp70s homologs (Small Angle Xray-Scattering and single-molecule Förster Resonance Energy Transfer) to which the MD simulations were compared in details. These structural ensembles represent the first attempt to model the conformational heterogeneity of a full-length molecular chaperone at the atomic scale. | ||
[[Image:JBSD30a.png|left|400px|thumb|<font color='blue'><b>NBD-IA is in blue</b></font>, <span style="color:deepskyblue;background-color:black;font-weight:bold;">NBD-IB is in deepskyblue</span>, <span style="color:lightsteelblue;background-color:black;font-weight:bold;">NBD-IIA is in lightsteelblue</span>, <span style="color:cyan;background-color:black;font-weight:bold;">NBD-IIB is in cyan</span>, <font color='magenta'><b>Linker is in magenta</b></font>, <span style="color:lime;background-color:black;font-weight:bold;">SBD-β is in green</span>, <font color='red'><b>SBD-α is in red</b></font>, <span style="color:darkgray;background-color:black;font-weight:bold;">C-terminal is in gray</span>]] | [[Image:JBSD30a.png|left|400px|thumb|<font color='blue'><b>NBD-IA is in blue</b></font>, <span style="color:deepskyblue;background-color:black;font-weight:bold;">NBD-IB is in deepskyblue</span>, <span style="color:lightsteelblue;background-color:black;font-weight:bold;">NBD-IIA is in lightsteelblue</span>, <span style="color:cyan;background-color:black;font-weight:bold;">NBD-IIB is in cyan</span>, <font color='magenta'><b>Linker is in magenta</b></font>, <span style="color:lime;background-color:black;font-weight:bold;">SBD-β is in green</span>, <font color='red'><b>SBD-α is in red</b></font>, <span style="color:darkgray;background-color:black;font-weight:bold;">C-terminal is in gray</span>]] | ||
| + | </StructureSection> | ||
== 3D Structures of Heat Shock Proteins == | == 3D Structures of Heat Shock Proteins == | ||
| Line 302: | Line 300: | ||
}} | }} | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
| + | <references/> | ||
Revision as of 12:14, 10 May 2015
| |||||||||||
3D Structures of Heat Shock Proteins
Updated on 10-May-2015
- ↑ Nicolai A, Delarue P, Senet P. Conformational dynamics of full-length inducible human Hsp70 derived from microsecond molecular dynamics simulations in explicit solvent. J Biomol Struct Dyn. 2012 Oct 17. PMID:23075261 doi:10.1080/07391102.2012.726190