Introduction:
GJB2 is a gene which encodes a member of the gap junction protein family. Intercellular signaling is one of the most essential properties of multicellular organisms. Gap junctions are specialized membrane regions containing hundreds of intercellular communication channels that allow the passage of molecules such as ions, metabolites, nucleotides and small peptides. The gap junctions were first characterized by electron microscopy as regionally specialized structures on plasma membranes of contacting adherent cells. These structures were shown to consist of cell-to-cell channels that facilitate the transfer of ions and small molecules between cells. The gap junction proteins, also known as connexins, purified from fractions of enriched gap junctions from different tissues differ. The gap junction proteins are divided into two categories, alpha and beta. Mutations in this gene are responsible for as much as 50% of pre-lingual, recessive deafness. [1]
How does mutation in connexin 26 make a new born deaf?
Mutations in human Connexin26 (hCx26) can lead to congenital hearing loss (1 child per 1000 frequency) that can be syndromic or non-syndromic. Non-syndromic hearing loss (NSHL) is characterized by sensorineural hearing loss in the absence of other symptoms, while syndromic hearing loss affects other organ systems, primarily the skin. mutations in GJB2 (the gene that encodes for Cx26) account for about half of all congenital and autosomal recessive nonsyndromic hearing loss in every population tested . Although the most frequently occurring NSHL mutations produce severely truncated proteins due to frameshift or missense, almost 80% of the known deafness mutations are actually single amino acid changes or deletions. These mutations have been found across the entire sequence of Cx26. The majority of NSHL mutations cause either generalized folding problems that result in the failure of Cx26 to traffic to the cell surface, or are permissive for the formation of gap junction plaques, but prevent intercellular channel function.[2]
Connexin26 (CX26) protein is essential for maintaining the high K+ concentration in the endolymph of the inner ear. Sound stimulation of the ossicular chain causes vibrations in the endolymph .K+ ions enter the hair cells under the influence of these vibrations and vibration signal is ultimately converted into a neural signal. The system is regenerated by the release of K+ from the hair cells into the supporting cells. The K+ ions are then passed from cell to cell via gap junctions and are eventually released into the endolymph. Except for sensorineural cells, the CX26 protein is present in gap junctions connecting all cell types in the cochlea , including the spiral limbus, the supporting cells, the spiral ligament and the basal and the basal and intermediate cells of the stria vascularis. It is therefore very likely that connexin 26 is involved in K+ -recycling in the cochlea.
Structure:
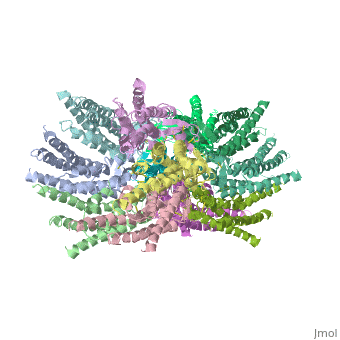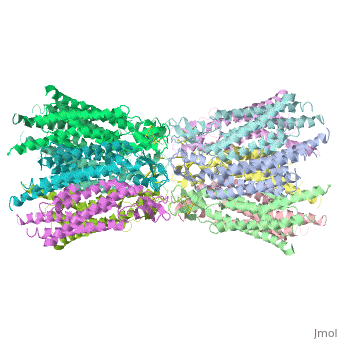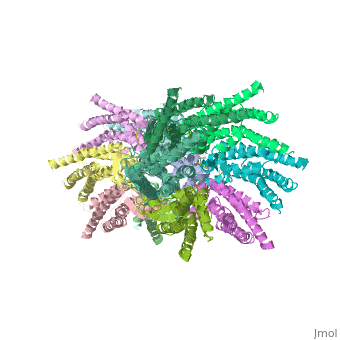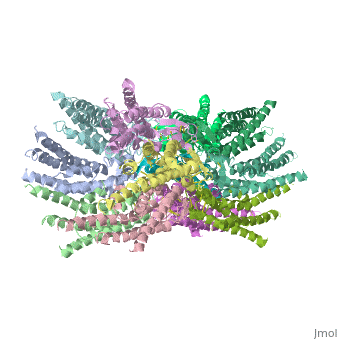
The overall structure of the Cx26 gap junction channel, which is formed by two connexons related to each other by a crystallographic two-fold symmetry axis, (Fig. 2a). It is a tsuzumi shape, a traditional Japanese drum. The protomers in each hexameric connexon are related by a sixfold non-crystallographic symmetry (NCS) axis perpendicular to the membrane plane (Fig. 2b). The height of the modelled structure of the gap junction channel without disordered cytoplasmic loop and C-terminal segment is approximately 155A °. The transmembrane region and membrane surfaces were deduced from the distribution of hydrophobic and aromatic amino acid residues along the noncrystallographic six-fold axis (Fig. 2a). The transmembrane region of the channel is 38A ° thick.TM2 extends about 19A ° from the membrane surface into the cytoplasm. The extracellular region of the connexon extends 23A ° from the membrane surface and interdigitates to the opposite connexon by 6A °, resulting in the intercellular ‘gap’ of 40A °. The extracellular lobes are not protruding so much, as indicated by the structural analyses of split gap junction channels with atomic force microscopy and electron microscopy. The relatively flat lobes could be attributed to the conformational change of the extracellular region induced by the docking of two connexons. The diameter of the connexon is biggest at the cytoplasmic side
of the membrane, 92A ° , and smallest at the extracellular side, 51A ° .
Viewed from the top, the channel looks like a ‘hexagonal nut’ with a pore in the centre (Fig. 2b).The diameter of the pore is about 40A ° at the cytoplasmic side of the channel, narrowing to 14A ° near the extracellular membrane surface and then widening to 25A ° in the extracellular space. [1]
Structure of the cx26 protomer:
The protomer has four transmembrane segments (TM1–4), two extracellular loops (E1 and E2), a cytoplasmic loop, an N-terminal helix (NTH), and a C-terminal segment (Fig. 3). Cx26 forms a typical four-helix bundle in which any pair of adjacent helices is antiparallel. TM1 and TM2 face the interior, whereas TM3 and TM4 face the hydrophobic membrane environment. There has been controversy about the identity of the major pore-lining helix, on the basis of accessibility studies of substituted cysteines and sequence analysis. One set of data favours TM3 as the major pore helix and the other favours TM1 . The helical arrangement of our structure is consistent with the latter model. The major pore-lining helix TM1 is inclined, so that the pore diameter narrows from the cytoplasmic to the extracellular side of the membrane, and ends in a short 310 helix.
The extracellular loop E1 contains a 310 helix at the beginning and a short a-helix in its C-terminal half E2, together with E1, contains a short antiparallel b-sheet and
stretches over E1, forming the outside wall of the connexon. Six conserved cysteine residues, three in each loop, form intramolecular disulphide bonds between E1 and E2 Most of the prominent intra-protomer interactions are in the extracellular part of the transmembrane region, Our structure revealed the interactions between the two adjoining connexons of the gap junction channel, which involve both E1 and E2 . The N-terminal half of E2 seems rather flexible
and its amino-acid sequence varies greatly among connexins . The C-terminal half of E2 begins with a 310 turn and is followed by a conserved Pro-Cys-Pro motif that reverses its direction back to TM4. Most of the prominent intra-protomer interactions are in the extracellular part of the transmembrane region (Fig. 4a ). Arg 32 (TM1) interactswithGln 80 (TM2),Glu 147 (TM3), and Ser 199 (TM4). Two hydrophobic cores around Trp 44 (E1) and Trp 77 (TM2) stabilize the protomer structure. Ala 39 (TM1), Ala 40 (TM1), Val 43 (E1) and Ile 74 (TM2) contribute to the first hydrophobic core around Trp 44, and Phe 154 (TM3) and Met 195 (TM4) form the second core with Trp 77 . In the intracellular part of the transmembrane region, Arg 143 (TM3) forms hydrogen bonds with Asn 206 (TM3) and Ser 139 (TM3) .[1]
Pore funnel and the voltage-dependent gating mechanism:
The short NTHs of the six protomers formthe funnel , This finding agrees with an NMR solution structure of an N-terminal peptide of Cx26, which showed that the loop connecting the NTH to TM1 is very flexible30. Asp 2 forms hydrogen bonds with the mainchain amide of Thr 5 from the neighbouring protomer. The Asp 2 and Thr 5 residues on neighbouring NTHs at the bottom of the funnel form a circular girdle, as previously seen in the nicotinic acetylcholine receptor31, which stabilizes the funnel structure . [1]
Differences between wild type and muatant connexin:
In general, single site mutations are spread fairly evenly across the whole protein with TM2 having the highest mutation density (number of amino acids with NHLS mutations divided by the total number of amino acids in the domain) at 67% to M1 and E1 having the lowest density of mutations with their respective domains at 33%. According to this criterion, TM4 has a mutation density of 40%. . Of the four transmembrane helices, M1, M2 and M3 have attracted the most attention, because of the controversies involved in models with different helix assignments, based on lower resolution cryo-electron crystallographic structures and scanning cysteine accessibility mutagenesis . Far less is known about TM4 and how side chains interact with the other helices and with the lipid bilayer. [2]
[3]
Structural highlights