We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
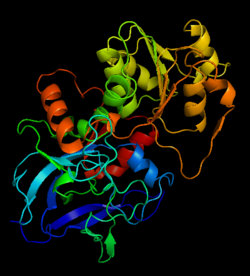
Alcohol dehydrogenase
From Proteopedia
(Difference between revisions)
| Line 74: | Line 74: | ||
{{Clear}} | {{Clear}} | ||
| - | The 3D structure of CbADH with the substitution Q100P (<scene name='2b83/Tet/3'>tetramer</scene>) was solved at 2.25 Å resolution ([[2b83]]). The <scene name='2b83/Mut/1'>substitution</scene> of Gln100 with Pro did not cause significant structural changes in the protein structure. The residues of the <span style="color:lime;background-color:black;font-weight:bold;">wildtype protein are colored green</ | + | The 3D structure of CbADH with the substitution Q100P (<scene name='2b83/Tet/3'>tetramer</scene>) was solved at 2.25 Å resolution ([[2b83]]). The <scene name='2b83/Mut/1'>substitution</scene> of Gln100 with Pro did not cause significant structural changes in the protein structure. The residues of the <span style="color:lime;background-color:black;font-weight:bold;">wildtype protein are colored green</span> and the residues of the <font color='cyan'><b>mutant one in cyan</b></font>. Only [http://en.wikipedia.org/wiki/Hydrogen_bond 2 H-bonds] were lost, one between Oε1 of Gln100 and the main chain N of Gly297, and the second between Nε2 of Gln100 and the main chain carbonyl O of Gly297. The mutation caused that an additional CH<sub>2</sub> group (Cδ of Pro100) is surrounded by nonpolar residues: Pro88 (3.8 Å), Trp90 (3.5 Å), and Val95 (4 Å). These residues (P100, P88, W90, and V95) are situated on a protruding lobe of the protein. An additional 11 [http://en.wikipedia.org/wiki/Aliphatic_compound aliphatic] and [http://en.wikipedia.org/wiki/Aromatic aromatic] carbon atoms are situated within the distance of 6 Å from Cδ of Pro100 (two [http://en.wikipedia.org/wiki/Methyl_group methyl groups] of Val95; three carbon atoms of the Trp90 [http://en.wikipedia.org/wiki/Indole indole] group; Cβ and Cγ [http://en.wikipedia.org/wiki/Methylene methylene] groups of Pro100; Cβ and Cγ of Gln101, and two carbons of the Phe99 [http://en.wikipedia.org/wiki/Phenyl_group phenyl] ring). |
{{Clear}} | {{Clear}} | ||
Revision as of 13:36, 18 May 2015
| |||||||||||
Additional Resources
For additional information, see: Carbohydrate Metabolism
3D Structures of Alcohol dehydrogenase
Updated on 18-May-2015
References
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol dehydrogenase from Human (Homo sapiens), different isozymes. SCOP. 2009. 1 March 2010 < http://scop.berkeley.edu/data/scop.b.d.c.b.b.c.html>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Bille V, Remacle J. Simple-kinetic descriptions of alcohol dehydrogenase after immobilization on tresyl-chloride-activated agarose. Eur J Biochem. 1986 Oct 15;160(2):343-8. PMID:3769934
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Blomstrand R, Ostling-Wintzell H, Lof A, McMartin K, Tolf BR, Hedstrom KG. Pyrazoles as inhibitors of alcohol oxidation and as important tools in alcohol research: an approach to therapy against methanol poisoning. Proc Natl Acad Sci U S A. 1979 Jul;76(7):3499-503. PMID:115004
- ↑ Alcohol Dehydrogenase. Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Alcohol Dehydrogenase.Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Goihberg E, Dym O, Tel-Or S, Levin I, Peretz M, Burstein Y. A single proline substitution is critical for the thermostabilization of Clostridium beijerinckii alcohol dehydrogenase. Proteins. 2007 Jan 1;66(1):196-204. PMID:17063493 doi:10.1002/prot.21170
- ↑ Goihberg E, Dym O, Tel-Or S, Shimon L, Frolow F, Peretz M, Burstein Y. Thermal stabilization of the protozoan Entamoeba histolytica alcohol dehydrogenase by a single proline substitution. Proteins. 2008 Feb 7;. PMID:18260103 doi:10.1002/prot.21946
- ↑ Goihberg E, Peretz M, Tel-Or S, Dym O, Shimon L, Frolow F, Burstein Y. Biochemical and Structural Properties of Chimeras Constructed by Exchange of Cofactor-Binding Domains in Alcohol Dehydrogenases from Thermophilic and Mesophilic Microorganisms. Biochemistry. 2010 Feb 9. PMID:20102159 doi:10.1021/bi901730x
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, David Birrer