This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1176
From Proteopedia
(Difference between revisions)
| Line 44: | Line 44: | ||
===Active State=== | ===Active State=== | ||
| - | After determining NTSR1-GW5 as only active-like, research was conducted to determine the structure of NTSR1 with catalytic nucleotide exchange. In order to do so, three of the six mutations were reverted back. The reversion of E166A, L310A, and F358A led to NTSR1 with G-protein activity at almost wild-type level. This protein was named, <scene name='72/721548/Ntsr1-elf/1'>NTSR1-ELF</scene>, and indicated that the amino acid residues E166, L310, and F358 play significant roles in the activity of NTSR1. | + | After determining NTSR1-GW5 as only active-like, research was conducted to determine the structure of NTSR1 with catalytic nucleotide exchange. In order to do so, three of the six mutations were reverted back, and these three residues were selected on the basis of their location. The reversion of E166A, L310A, and F358A led to NTSR1 with G-protein activity at almost wild-type level. This protein was named, <scene name='72/721548/Ntsr1-elf/1'>NTSR1-ELF</scene>, and indicated that the amino acid residues E166, L310, and F358 play significant roles in the activity of NTSR1. |
====L310==== | ====L310==== | ||
| Line 56: | Line 56: | ||
====E166==== | ====E166==== | ||
| - | Although the role of E166 in G-protein activity is not quite as clear as it is for L310 or F358, mutating this residue to an alanine significantly reduced catalytic nucleotide exchange. E166 is part of a D/ERY motif that is highly conserved in class A GPCRs. This motif includes R167, which is positioned by L310, as mentioned. | + | Although the role of E166 in G-protein activity is not quite as clear as it is for L310 or F358, mutating this residue to an alanine significantly reduced catalytic nucleotide exchange. E166 is part of a D/ERY motif that is highly conserved in class A GPCRs. This motif includes R167, which is positioned by L310, as mentioned. In order to hypothesize a role for E166, Krumm et al looked at other class A GPCRs. In the active M2 receptor, the acidic amino acid is stabilized by a hydrogen bond with N58, and this N58 plays a direct role in stabilization of the G protein. The interaction of E166 in NTSR1 has a similar interaction with V102 and T101, along with a weak interaction with H105. The researchers then looked at the β2AR receptor and found Y141 positioned intracellular helix 2, and this interaction was linked with the D/ERY motif. This was determined an essential interaction for G-protein activity, and M181 may cause an equivalent interaction in NTSR1. E166 and the residues it is hypothesized to interact with are shown below. |
| + | |||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 01:14, 31 March 2016
| This Sandbox is Reserved from Jan 11 through August 12, 2016 for use in the course CH462 Central Metabolism taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1160 through Sandbox Reserved 1184. |
To get started:
More help: Help:Editing |
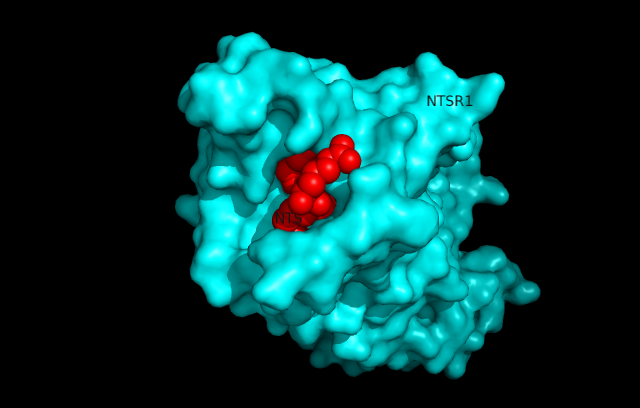
Rattus norevegicus NTSR1
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644