This is a default text for your page Allison Cotter/Sandbox 1. Click above on edit this page to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Introduction
Neurotensin receptor 1 (NTSR1) is a G-protein coupled receptor (GPCR) that binds to the 13 amino acid peptide, neurotensin. Studies determining the structure of NTSR1 crystallized the GPCR bound with the C-terminus of its tridecapeptide ligand, because it has a higher potency and efficacy than its full-length counterpart. NTSR1 is a class A GPCR, and like all G-proteins, consists of an extracellular binding domain along with 7 transmembrane helices. Along with the ligand binding pocket at the top of the protein, NTSR1 also contains an allosteric Na+ ion binding pocket underneath. NTS binds to NTSR1, leading to a conformational change of the protein and modulation of second messengers. NTS has been shown to have a variety of biological activities including a role in the leptin signalling pathways, tumor growth, and dopamine regulation. The majority of effects of NTS are mediated through NTSR1. Research of the structure of NTSR1 has focused on the differences between its active and active-like states.
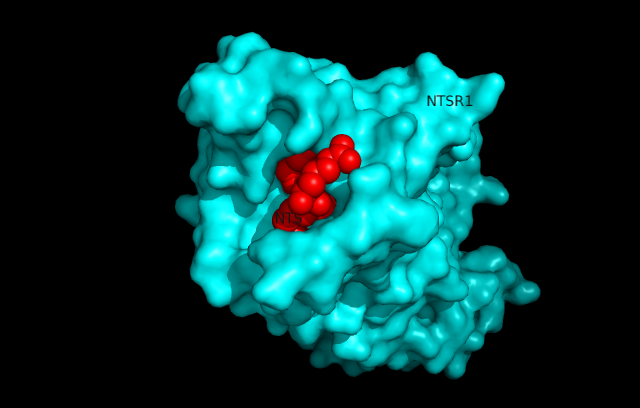
Structure
Ligand Binding Pocket
Near the top of the protein is the
This pocket is comprised of several key amino acid resides. The first reside is a Phenylalanine at position F358. The purpose of this amino acid is to take part in a network of hydrophobic stacking interactions. These interactions stabilize the Y324 and W321 residues. These are crucial interactions because the Y324 is the amino acid residue that directly interacts with the
via https://en.wikipedia.org/wiki/Van_der_Waals_force Van der Waals interactions.
Without the hydrophobic stacking interactions that are facilitated by the F358 this binding interaction would not occur. The W321 residue also partakes in these stacking interactions. The W321 serves as the boundary between the ligand binding pocket and the sodium binding pocket.
Na+ Binding Pocket
, which is positioned at the bottom of the , sets the top of the . The Na+ ion binding pocket acts as a negative allosteric site for G protein activity. When Na+ enters the Na+ ion binding pocket, it coordinates with Asp95, Gln131, and S135, and shuts down the activity of the protein. When the G protein is in its active state, the Na+ ion binding pocket is collapsed, preventing the regulation of protein activity through a Na+ ion. In this case, the Na+ ion is coordinated by a salt bridge to Asp113. The side chain atoms of Asp113 form a hydrogen bond network with Thr156, Ser361, Ser362, and Gln365, which prevents the coordination of a Na+ ion.
Neurotensin
is a 13 amino acid peptide that is found in both nervous and peripheral tissues. It functions as a hormone and a neurotransmitter by activating as the ligand for the G-protein coupled receptor NTSR1.
Biological Relevance
Leptin Research
A study was done by Liang Y. et al.to analyze the effects that being NTSR1 deficient had on mice. The results demonstrated that mice who were NTSR1 deficient were not able to receive a satiety signal.This resulted in the mice to continuing to eat as long as food was present, leading to significant weight gain. This is due to the fact that NTSR1 is involved in a signaling pathway that regulates leptin and therefore food intake. Without sufficient NTSR1 this pathway is interrupted. https://en.wikipedia.org/wiki/Leptin
Cancer Studies
It has been shown that some tumor cells can secrete and express Neurotensin and Neurotensin receptors themselves suggetsing that autocrine, endocrine and paracrine regulation by Neurotensin are possible. This leads to aggressive growth and possibly tumor development. A study was done that demonstrated that injecting animals with Neurotensin increased tumor growth and size, while injecting them with Neurotensin antagonist decreased tumor growth.It is believed that Neurotensin regulation may be used in future cancer treatment techniques.
Schizophrenia Research
The Dopamine Hypothesis http://www.schizophreniaforum.org/for/curr/AbiDargham/ states that having hyperdopamine levels may lead to schizophrenic symptoms. It has been shown that NTSR1 caused a blockade which inhibited firing in dopaminergic cells. It is believed that NTSR1 could therefore be used as a therapeutic for treating schizophrenia.Although this research is promising, the secondary effects were too extreme and the trial was discontinued.This is a pathway where NTSR1 research is focused on that could lead to ground breaking advances in treating schizophrenia.
Structural Research
Studies have been done to decipher the structure of NTSR1. Since wild type NTSR1 is unstable in detergent solution, select residues in the protein were mutated for stabilization. None of these residues are in the NTS binding pocket.
Active-Like State
Originally, six different amino acids were mutated for thermostabilization in detergent solution. These mutations were A86L, E166A, G215A, L310A, F358A, and V360A. This protein was found to have NTS affinity similar to that of wild tpye NTSR1, and was named . Along with this, the Na+ ion binding pocket was found to be collapsed in this protein. However, studies found that NTSR1-GW5 did not have G-protein activity.
Active State
After determining as only active-like, research was conducted to determine the structure of NTSR1 with catalytic nucleotide exchange. In order to do so, three of the six mutations were reverted back, and these three residues were selected on the basis of their location. The reversion of E166A, L310A, and F358A led to NTSR1 with G-protein activity at almost wild-type level. This protein was named, , and indicated that the amino acid residues E166, L310, and F358 play significant roles in the activity of NTSR1.
L310
F358
E166
Although the role of E166 in G-protein activity is not quite as clear as it is for L310 or F358, mutating this residue to an alanine significantly reduced catalytic nucleotide exchange. E166 is part of a D/ERY motif that is highly conserved in class A GPCRs. This motif includes R167, which is positioned by L310, as mentioned. In order to hypothesize a role for E166, Krumm et al looked at other class A GPCRs. In the active M2 receptor, the acidic amino acid is stabilized by a hydrogen bond with N58, and this N58 plays a direct role in stabilization of the G protein. The interaction of E166 in NTSR1 has a similar interaction with V102 and T101, along with a weak interaction with H105. The researchers then looked at the β2AR receptor and found Y141 positioned intracellular helix 2, and this interaction was linked with the D/ERY motif. This was determined an essential interaction for G-protein activity, and M181 may cause an equivalent interaction in NTSR1. E166 and the residues it is hypothesized to interact with are shown below.
Image:E166 for Proteopedia page.png

