This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
TEM1 Class Antibiotic Resistance Proteins
From Proteopedia
(Difference between revisions)
| Line 11: | Line 11: | ||
<tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat">[[1axb|1axb]], [[1bt5|1bt5]], [[1btl|1btl]], [[1ck3|1ck3]], [[1erm|1erm]], [[1ero|1ero]], [[1erq|1erq]], [[1esu|1esu]], [[1fqg|1fqg]], [[1jtd|1jtd]], [[1jtg|1jtg]], [[1jvj|1jvj]], [[1jwp|1jwp]], [[1jwv|1jwv]], [[1jwz|1jwz]], [[1lhy|1lhy]], [[1li0|1li0]], [[1li9|1li9]], [[1m40|1m40]], [[1nxy|1nxy]], [[1ny0|1ny0]], [[1nym|1nym]], [[1nyy|1nyy]], [[1pzo|1pzo]], [[1pzp|1pzp]], [[1s0w|1s0w]], [[1tem|1tem]], [[1xxm|1xxm]], [[1yt4|1yt4]], [[1zg4|1zg4]], [[1zg6|1zg6]], [[2v1z|2v1z]]</td></tr> | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat">[[1axb|1axb]], [[1bt5|1bt5]], [[1btl|1btl]], [[1ck3|1ck3]], [[1erm|1erm]], [[1ero|1ero]], [[1erq|1erq]], [[1esu|1esu]], [[1fqg|1fqg]], [[1jtd|1jtd]], [[1jtg|1jtg]], [[1jvj|1jvj]], [[1jwp|1jwp]], [[1jwv|1jwv]], [[1jwz|1jwz]], [[1lhy|1lhy]], [[1li0|1li0]], [[1li9|1li9]], [[1m40|1m40]], [[1nxy|1nxy]], [[1ny0|1ny0]], [[1nym|1nym]], [[1nyy|1nyy]], [[1pzo|1pzo]], [[1pzp|1pzp]], [[1s0w|1s0w]], [[1tem|1tem]], [[1xxm|1xxm]], [[1yt4|1yt4]], [[1zg4|1zg4]], [[1zg6|1zg6]], [[2v1z|2v1z]]</td></tr> | ||
</table> | </table> | ||
| + | |||
| + | == Background and History == | ||
| + | Antibiotics have long been the primary line of defense against many infections. The advent of penicillin in the 1930s brought an end to an era of uncertainty: diseases once considered fatal became treatable. Unfortunately, natural selection stops for no species. Resistance to some form of antibiotic now has become standard for many infections.[A] Moreover, their liberal usage have resulted in “superbugs”, which are resistant to multiple antibiotics.[B] These strains are capable of secreting enzymes capable of deactivating the antibiotic or modifying their cell walls to render the antibiotic ineffective. | ||
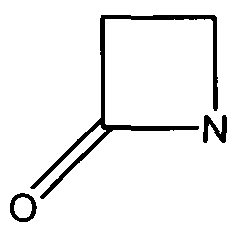
| + | Many antibiotics such as penicillin, their derivatives (penams), and cephalosporins (cephams) possess a β-lactam ring (Figure 1). | ||
| + | [[Image:Beta-Lactam-Ring.png]] | ||
| + | Figure 1. The beta-lactam ring is highlighted in red. Beta-lactamases function in hydrolyzing the amide bond within the ring, rendering the antibiotic ineffective. (Image Credit: Wikipedia) | ||
| + | |||
| + | β-Lactamases function by hydrolyzing the β-lactam ring within the antibiotic (Figure 2). This prevents the interaction between the cell wall and the antibiotic. | ||
| + | [[Image:Figure2.jpeg]] | ||
| + | Figure 2. Bacteria are capable of becoming antibiotic resistant by catalyzing the hydrolysis of the β-lactam ring. | ||
| + | |||
| + | The TEM-1 subclass is one of the most common of all β-Lactamases.[3] Their prevalence began when bacteria started exhibiting penicillin resistance on a mass scale. Since the TEM-1 subclass has been prevalent for many years, many inhibitors have either been discovered or synthesized to prevent their catalytic function. | ||
== Function == | == Function == | ||
Revision as of 22:46, 13 April 2016
Your Heading Here (maybe something like 'Structure')
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
Proteopedia Page Contributors and Editors (what is this?)
Kenna Salvatore, Matt O'Malley, Ryan Hunter Wilson, Riley Culhane, Michal Harel