This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1165
From Proteopedia
(Difference between revisions)
| Line 27: | Line 27: | ||
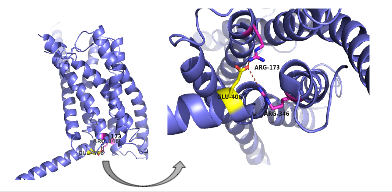
| - | There are specific amino acid interactions that hold the helices of the 7TM in the conformation that maximizes [http://www.chemicool.com/definition/affinity.html affinity]. <ref name="Ligands">PMID: 21542831</ref> | + | There are specific amino acid interactions that maximize affinity. This includes the alpha helical structure of the <scene name='72/721535/Opening_orientation/2'>stalk</scene>. The alpha helical structure of the stalk interacts directly with glucagon; when the alpha helix of the stalk is disrupted, the affinity of glucagon for GCGR decreases. Furthermore, there are certain interactions that hold the helices of the 7TM in the conformation that maximizes [http://www.chemicool.com/definition/affinity.html affinity]. <ref name="Ligands">PMID: 21542831</ref> The high affinity conformation of GCGR is the open conformation when glucagon can bind. Without these specific interactions between the residues, open conformation is not stabilized and GCGR remains in the closed conformation where glucagon cannot bind. <ref name="Tips">PMID: 23863937</ref> The [https://en.wikipedia.org/wiki/Disulfide disulfide bond] between <scene name='72/721535/Disulfide_bond_notspin/1'> Cys 294 and Cys 224</scene> serves to hold the helices in the proper orientation for binding and stabilize the open conformation. Additionally, the [https://en.wikipedia.org/wiki/Salt_bridge_%28protein_and_supramolecular%29 salt bridges] between Glu 406, Arg 173, and Arg 346 hold the open conformation together for higher affinity (Figure 4). <ref name="Ligands">PMID: 21542831</ref> |
| - | [[Image:Screen Shot 2016-03-29 at 3.24.43 PM.png|(|):|400 px|center|thumb|'''Figure | + | [[Image:Screen Shot 2016-03-29 at 3.24.43 PM.png|(|):|400 px|center|thumb|'''Figure 4: Salt Bridge'''. The non-covalent interactions between residues Glu 406, Arg 173, and Arg 346 form a [https://en.wikipedia.org/wiki/Denticity tridentate] salt bridge. The Glu 406 acts as the central residue in the tridentate salt bridge; Arg 173 and Arg 436 both interact with Glu 406. The salt bridge is located on the intracellular side of the transmembrane helices.]] |
Revision as of 23:34, 18 April 2016
References
- ↑ 1.0 1.1 Hollenstein K, de Graaf C, Bortolato A, Wang MW, Marshall FH, Stevens RC. Insights into the structure of class B GPCRs. Trends Pharmacol Sci. 2014 Jan;35(1):12-22. doi: 10.1016/j.tips.2013.11.001. Epub, 2013 Dec 18. PMID:24359917 doi:http://dx.doi.org/10.1016/j.tips.2013.11.001
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 Siu FY, He M, de Graaf C, Han GW, Yang D, Zhang Z, Zhou C, Xu Q, Wacker D, Joseph JS, Liu W, Lau J, Cherezov V, Katritch V, Wang MW, Stevens RC. Structure of the human glucagon class B G-protein-coupled receptor. Nature. 2013 Jul 25;499(7459):444-9. doi: 10.1038/nature12393. Epub 2013 Jul 17. PMID:23863937 doi:10.1038/nature12393
- ↑ 3.0 3.1 3.2 3.3 3.4 Miller LJ, Dong M, Harikumar KG. Ligand binding and activation of the secretin receptor, a prototypic family B G protein-coupled receptor. Br J Pharmacol. 2012 May;166(1):18-26. doi: 10.1111/j.1476-5381.2011.01463.x. PMID:21542831 doi:http://dx.doi.org/10.1111/j.1476-5381.2011.01463.x
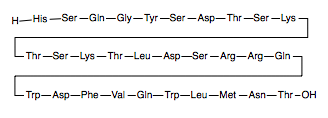
- ↑ Thomsen J, Kristiansen K, Brunfeldt K, Sundby F. The amino acid sequence of human glucagon. FEBS Lett. 1972 Apr 1;21(3):315-319. PMID:11946536
- ↑ 5.0 5.1 5.2 Bortolato A, Dore AS, Hollenstein K, Tehan BG, Mason JS, Marshall FH. Structure of Class B GPCRs: new horizons for drug discovery. Br J Pharmacol. 2014 Jul;171(13):3132-45. doi: 10.1111/bph.12689. PMID:24628305 doi:http://dx.doi.org/10.1111/bph.12689
- ↑ Mukund S, Shang Y, Clarke HJ, Madjidi A, Corn JE, Kates L, Kolumam G, Chiang V, Luis E, Murray J, Zhang Y, Hotzel I, Koth CM, Allan BB. Inhibitory mechanism of an allosteric antibody targeting the glucagon receptor. J Biol Chem. 2013 Nov 4. PMID:24189067 doi:http://dx.doi.org/10.1074/jbc.M113.496984
- ↑ Hoare SR. Allosteric modulators of class B G-protein-coupled receptors. Curr Neuropharmacol. 2007 Sep;5(3):168-79. doi: 10.2174/157015907781695928. PMID:19305799 doi:http://dx.doi.org/10.2174/157015907781695928
- ↑ 8.0 8.1 8.2 Yang L, Yang D, de Graaf C, Moeller A, West GM, Dharmarajan V, Wang C, Siu FY, Song G, Reedtz-Runge S, Pascal BD, Wu B, Potter CS, Zhou H, Griffin PR, Carragher B, Yang H, Wang MW, Stevens RC, Jiang H. Conformational states of the full-length glucagon receptor. Nat Commun. 2015 Jul 31;6:7859. doi: 10.1038/ncomms8859. PMID:26227798 doi:http://dx.doi.org/10.1038/ncomms8859