This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1p7q
From Proteopedia
| Line 7: | Line 7: | ||
|ACTIVITY= | |ACTIVITY= | ||
|GENE= HLA-A OR HLAA ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]), B2M ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]) | |GENE= HLA-A OR HLAA ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]), B2M ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]) | ||
| + | |DOMAIN= | ||
| + | |RELATEDENTRY= | ||
| + | |RESOURCES=<span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1p7q FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1p7q OCA], [http://www.ebi.ac.uk/pdbsum/1p7q PDBsum], [http://www.rcsb.org/pdb/explore.do?structureId=1p7q RCSB]</span> | ||
}} | }} | ||
| Line 16: | Line 19: | ||
==Disease== | ==Disease== | ||
| - | Known | + | Known disease associated with this structure: Hypoproteinemia, hypercatabolic OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=109700 109700]] |
==About this Structure== | ==About this Structure== | ||
| Line 30: | Line 33: | ||
[[Category: hla-a2/lir-1 complex]] | [[Category: hla-a2/lir-1 complex]] | ||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Sun Mar 30 22:56:03 2008'' |
Revision as of 19:56, 30 March 2008
| |||||||
| , resolution 3.40Å | |||||||
|---|---|---|---|---|---|---|---|
| Gene: | HLA-A OR HLAA (Homo sapiens), B2M (Homo sapiens) | ||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
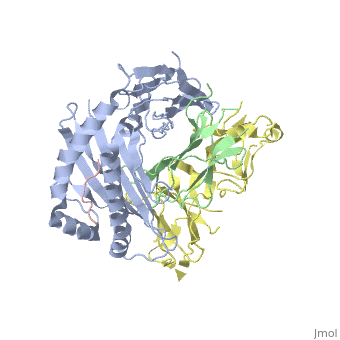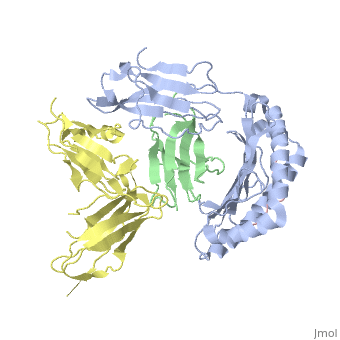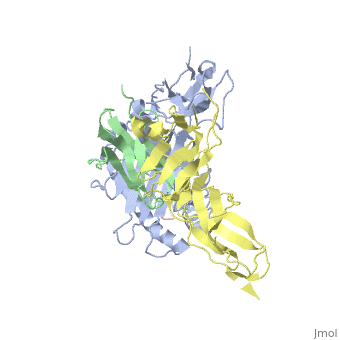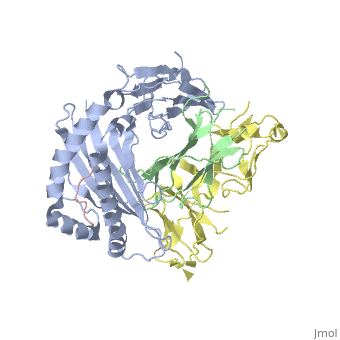
Crystal Structure of HLA-A2 Bound to LIR-1, a Host and Viral MHC Receptor
Contents |
Overview
Leukocyte immunoglobulin-like receptor 1 (LIR-1), an inhibitory receptor expressed on monocytes, dendritic cells and lymphocytes, regulates cellular function by binding a broad range of classical and nonclassical major histocompatibility complex (MHC) class I molecules, and the human cytomegalovirus MHC class I homolog UL18. Here we describe the 3.4-A crystal structure of a complex between the LIR-1 D1D2 domains and the MHC class I molecule HLA-A2. LIR-1 contacts the mostly conserved beta(2)-microglobulin and alpha3 domains of HLA-A2. The LIR-1 binding site comprises residues at the interdomain hinge, and a patch at the D1 tip. The structure shows how LIR-1 recognizes UL18 and diverse MHC class I molecules, and indicates that a similar mode of MHC class I recognition is used by other LIR family members.
Disease
Known disease associated with this structure: Hypoproteinemia, hypercatabolic OMIM:[109700]
About this Structure
1P7Q is a Protein complex structure of sequences from Homo sapiens. Full crystallographic information is available from OCA.
Reference
Crystal structure of HLA-A2 bound to LIR-1, a host and viral major histocompatibility complex receptor., Willcox BE, Thomas LM, Bjorkman PJ, Nat Immunol. 2003 Sep;4(9):913-9. Epub 2003 Aug 3. PMID:12897781
Page seeded by OCA on Sun Mar 30 22:56:03 2008