We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
CRISPR-Cas
From Proteopedia
(Difference between revisions)
| Line 42: | Line 42: | ||
The 2 classes of CRISPR-Cas systems differ fundamentally with respect to the organization of the effector module <ref name="Rev430">doi:10.1038/nrmicro3569</ref>. Class 1 systems (including types I, III, and IV) are present in bacteria and archaea, and encompass effector complexes composed of 4-7 Cas protein subunits [''e.g.'', the ('''C'''RISPR-'''as'''sociated '''c'''omplex for '''a'''ntiviral '''de'''fense) ('''Cascade''') of type I systems, and the Csm/Cmr complexes of type III systems]. Most of the subunits of the class 1 effector complexes — in particular, Cas5, Cas6, and Cas7—contain variants of the RNA-binding RRM (RNA recognition motif) domain. Although the sequence similarity between the individual subunits of type I and type III effector complexes is generally low, the complexes share strikingly similar overall architectures that suggest a common origin <ref name="Rev431">doi:10.1016/j.molcel.2015.10.008</ref>. The ancestral CRISPR-Cas effector complex most likely resembled the extant type III complexes, as indicated by the presence of the archetypal type III protein, the large Cas10 subunit, which appears to be an active enzyme of the DNA polymerase–nucleotide cyclase superfamily, unlike its inactive type I counterpart (Cas8) <ref name="Rev431">doi:10.1016/j.molcel.2015.10.008</ref>. The ''cas6'' gene family encodes a set of RNA endonucleases responsible for crRNA processing in Type I and Type III CRISPR systems. Type II systems use a trans-activating RNA (tracrRNA) together with endogenous RNase III for crRNA maturation (Figure 2).In Type I-B, I-C, I-E, and I-F systems, the endoRNase stays bound to the crRNA and assembles into a complex with other Cas proteins for downstream targeting <ref name="Rev312">doi:10.1126/science.1159689</ref>, while in Type I-A and III systems, the crRNA alone is loaded into the targeting complex and Cas6 dissociates <ref name="Rev3">PMID:25468820</ref>. | The 2 classes of CRISPR-Cas systems differ fundamentally with respect to the organization of the effector module <ref name="Rev430">doi:10.1038/nrmicro3569</ref>. Class 1 systems (including types I, III, and IV) are present in bacteria and archaea, and encompass effector complexes composed of 4-7 Cas protein subunits [''e.g.'', the ('''C'''RISPR-'''as'''sociated '''c'''omplex for '''a'''ntiviral '''de'''fense) ('''Cascade''') of type I systems, and the Csm/Cmr complexes of type III systems]. Most of the subunits of the class 1 effector complexes — in particular, Cas5, Cas6, and Cas7—contain variants of the RNA-binding RRM (RNA recognition motif) domain. Although the sequence similarity between the individual subunits of type I and type III effector complexes is generally low, the complexes share strikingly similar overall architectures that suggest a common origin <ref name="Rev431">doi:10.1016/j.molcel.2015.10.008</ref>. The ancestral CRISPR-Cas effector complex most likely resembled the extant type III complexes, as indicated by the presence of the archetypal type III protein, the large Cas10 subunit, which appears to be an active enzyme of the DNA polymerase–nucleotide cyclase superfamily, unlike its inactive type I counterpart (Cas8) <ref name="Rev431">doi:10.1016/j.molcel.2015.10.008</ref>. The ''cas6'' gene family encodes a set of RNA endonucleases responsible for crRNA processing in Type I and Type III CRISPR systems. Type II systems use a trans-activating RNA (tracrRNA) together with endogenous RNase III for crRNA maturation (Figure 2).In Type I-B, I-C, I-E, and I-F systems, the endoRNase stays bound to the crRNA and assembles into a complex with other Cas proteins for downstream targeting <ref name="Rev312">doi:10.1126/science.1159689</ref>, while in Type I-A and III systems, the crRNA alone is loaded into the targeting complex and Cas6 dissociates <ref name="Rev3">PMID:25468820</ref>. | ||
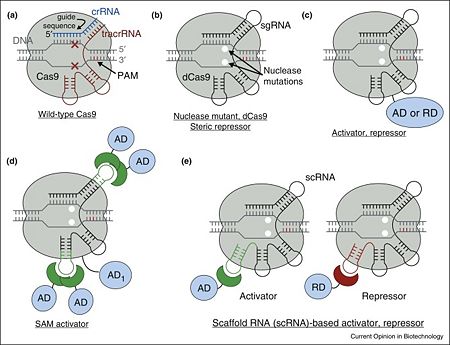
| - | In the less common class 2 CRISPR-Cas systems (types II, V, and VI), which are almost completely restricted to bacteria, the effector complex is represented by a single multidomain protein <ref name="Rev430">doi:10.1038/nrmicro3569</ref>. The best-characterized class 2 effector is Cas9 (type II), the RNA-dependent endonuclease that contains two unrelated nuclease domains, HNH and RuvC, that are responsible for the cleavage of the target and the displaced strand, respectively, in the crRNA–target DNA complex (<scene name='74/746096/Cv3/1'>Domain organization of nuclease lobe of Cas9 from S. pyogenes</scene>, [[4zt0]]) The type II loci also encode a trans-acting CRISPR RNA (tracrRNA) that evolved from the corresponding CRISPR repeat and is essential for pre-crRNA processing and target recognition in type II systems | + | In the less common class 2 CRISPR-Cas systems (types II, V, and VI), which are almost completely restricted to bacteria, the effector complex is represented by a single multidomain protein <ref name="Rev430">doi:10.1038/nrmicro3569</ref>. The best-characterized class 2 effector is Cas9 (type II), the RNA-dependent endonuclease that contains two unrelated nuclease domains, HNH and RuvC, that are responsible for the cleavage of the target and the displaced strand, respectively, in the crRNA–target DNA complex (<scene name='74/746096/Cv3/1'>Domain organization of nuclease lobe of Cas9 from S. pyogenes</scene>, [[4zt0]]). The type II loci also encode a trans-acting CRISPR RNA (tracrRNA) that evolved from the corresponding CRISPR repeat and is essential for pre-crRNA processing and target recognition in type II systems. Cas9 is directed to its DNA targets by forming a ribonucleoprotein complex with these 2 small non-coding RNAs: crRNA and tracrRNA. By elegant engineering, <scene name='74/742625/Cv3/8'>crRNA and tracrRNA can be joined end-to-end and transcribed as a single guide RNA (sgRNA)</scene> ([[4zt9]]<ref name="dCAS9">PMID:26113724</ref>) that too efficiently directs Cas9 protein to DNA targets encoded within the guide sequence of sgRNA <ref name="Jinek">PMID:22745249</ref>: |
| - | + | ||
| - | Cas9 is directed to its DNA targets by forming a ribonucleoprotein complex with | + | |
''Examples of 3D structures of single guide RNA (sgRNA)'' | ''Examples of 3D structures of single guide RNA (sgRNA)'' | ||
| Line 50: | Line 48: | ||
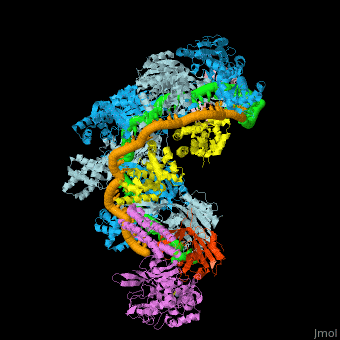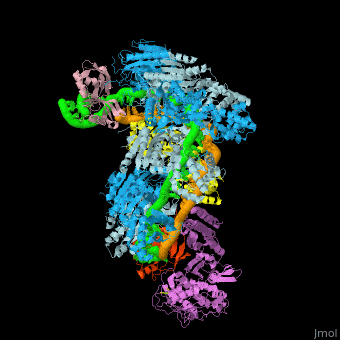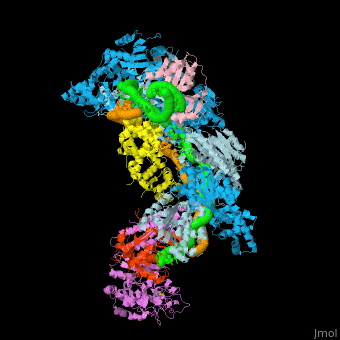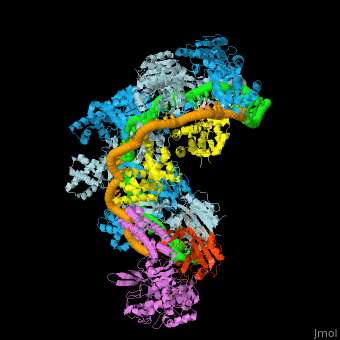
*<scene name='74/742625/Cv2/12'>Cas9-sgRNA-target DNA complex from Streptococcus pyogenes</scene> ([[5b2s]]). | *<scene name='74/742625/Cv2/12'>Cas9-sgRNA-target DNA complex from Streptococcus pyogenes</scene> ([[5b2s]]). | ||
*<scene name='74/742625/Cv2/13'>Cas9-sgRNA-target DNA complex from Francisella tularensis</scene> ([[5b2p]]). | *<scene name='74/742625/Cv2/13'>Cas9-sgRNA-target DNA complex from Francisella tularensis</scene> ([[5b2p]]). | ||
| - | *<scene name='74/742625/Cv3/2'>Cas9-sgRNA-target DNA complex from Staphylococcus aureus</scene> ([[4axw]]). | + | *<scene name='74/742625/Cv3/2'>Cas9-sgRNA-target DNA complex from Staphylococcus aureus</scene> ([[4axw]]). |
| + | |||
| + | The prototype type V effector Cpf1 (subtype V-A) contains only one nuclease domain (RuvC-like) that is identifiable by sequence analysis (39). However, analysis of the recently solved structure of Cpf1 complexed with the crRNA and target DNA has revealed a second nuclease domain, the fold of which is unrelated to HNH or any other known nucleases. In analogy to the HNH domain in Cas9, the novel nuclease domain in Cpf1 is inserted into the RuvC domain, and it is responsible for cleavage of the target strand (40). | ||
The <scene name='74/742625/Cv3/4'>optimal DNA target of the complex is determined by a Watson–Crick base pairing of a short ∼20-nt sequence within sgRNA (within the crRNA in wild-type)</scene>, termed the guide sequence, adjacent to a <scene name='74/742625/Cv3/10'>few nucleotide long conserved motif recognized directly by Cas9 protein (protospacer adjacent motif, PAM)</scene> <ref name="Jinek">PMID:22745249</ref><ref name="Prin6">PMID:22949671</ref>. Despite this, a <scene name='74/742625/Cv/44'>few mismatches between guide sequence and target DNA can be tolerated</scene> <ref name="Jinek">PMID:22745249</ref><ref name="Prin7">PMID:23452860</ref><ref name="Prin8">PMID:23761437</ref><ref name="Prin9">PMID:24837660</ref>, more so within the 5’ proximal position of the guide sequence. | The <scene name='74/742625/Cv3/4'>optimal DNA target of the complex is determined by a Watson–Crick base pairing of a short ∼20-nt sequence within sgRNA (within the crRNA in wild-type)</scene>, termed the guide sequence, adjacent to a <scene name='74/742625/Cv3/10'>few nucleotide long conserved motif recognized directly by Cas9 protein (protospacer adjacent motif, PAM)</scene> <ref name="Jinek">PMID:22745249</ref><ref name="Prin6">PMID:22949671</ref>. Despite this, a <scene name='74/742625/Cv/44'>few mismatches between guide sequence and target DNA can be tolerated</scene> <ref name="Jinek">PMID:22745249</ref><ref name="Prin7">PMID:23452860</ref><ref name="Prin8">PMID:23761437</ref><ref name="Prin9">PMID:24837660</ref>, more so within the 5’ proximal position of the guide sequence. | ||
Revision as of 14:05, 27 November 2016
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 Didovyk A, Borek B, Tsimring L, Hasty J. Transcriptional regulation with CRISPR-Cas9: principles, advances, and applications. Curr Opin Biotechnol. 2016 Aug;40:177-84. doi: 10.1016/j.copbio.2016.06.003. Epub, 2016 Jun 23. PMID:27344519 doi:http://dx.doi.org/10.1016/j.copbio.2016.06.003
- ↑ Brophy JA, Voigt CA. Principles of genetic circuit design. Nat Methods. 2014 May;11(5):508-20. doi: 10.1038/nmeth.2926. PMID:24781324 doi:http://dx.doi.org/10.1038/nmeth.2926
- ↑ Straubeta A, Lahaye T. Zinc fingers, TAL effectors, or Cas9-based DNA binding proteins: what's best for targeting desired genome loci? Mol Plant. 2013 Sep;6(5):1384-7. doi: 10.1093/mp/sst075. Epub 2013 May 29. PMID:23718948 doi:http://dx.doi.org/10.1093/mp/sst075
- ↑ Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol. 2014 Apr;32(4):347-55. doi: 10.1038/nbt.2842. Epub 2014 Mar 2. PMID:24584096 doi:http://dx.doi.org/10.1038/nbt.2842
- ↑ 5.0 5.1 5.2 5.3 5.4 5.5 5.6 Hochstrasser ML, Doudna JA. Cutting it close: CRISPR-associated endoribonuclease structure and function. Trends Biochem Sci. 2015 Jan;40(1):58-66. doi: 10.1016/j.tibs.2014.10.007. Epub, 2014 Nov 18. PMID:25468820 doi:http://dx.doi.org/10.1016/j.tibs.2014.10.007
- ↑ 6.0 6.1 Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007 Mar 23;315(5819):1709-12. PMID:17379808 doi:http://dx.doi.org/10.1126/science.1138140
- ↑ 7.00 7.01 7.02 7.03 7.04 7.05 7.06 7.07 7.08 7.09 7.10 Mohanraju P, Makarova KS, Zetsche B, Zhang F, Koonin EV, van der Oost J. Diverse evolutionary roots and mechanistic variations of the CRISPR-Cas systems. Science. 2016 Aug 5;353(6299):aad5147. doi: 10.1126/science.aad5147. PMID:27493190 doi:http://dx.doi.org/10.1126/science.aad5147
- ↑ Kunin V, Sorek R, Hugenholtz P. Evolutionary conservation of sequence and secondary structures in CRISPR repeats. Genome Biol. 2007;8(4):R61. PMID:17442114 doi:http://dx.doi.org/10.1186/gb-2007-8-4-r61
- ↑ 9.0 9.1 9.2 Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, van der Oost J. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science. 2008 Aug 15;321(5891):960-4. doi: 10.1126/science.1159689. PMID:18703739 doi:http://dx.doi.org/10.1126/science.1159689
- ↑ Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadan AH, Moineau S. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature. 2010 Nov 4;468(7320):67-71. doi: 10.1038/nature09523. PMID:21048762 doi:http://dx.doi.org/10.1038/nature09523
- ↑ 11.0 11.1 11.2 11.3 11.4 Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ, Barrangou R, Brouns SJ, Charpentier E, Haft DH, Horvath P, Moineau S, Mojica FJ, Terns RM, Terns MP, White MF, Yakunin AF, Garrett RA, van der Oost J, Backofen R, Koonin EV. An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol. 2015 Nov;13(11):722-36. doi: 10.1038/nrmicro3569. Epub 2015, Sep 28. PMID:26411297 doi:http://dx.doi.org/10.1038/nrmicro3569
- ↑ 12.0 12.1 12.2 12.3 Shmakov S, Abudayyeh OO, Makarova KS, Wolf YI, Gootenberg JS, Semenova E, Minakhin L, Joung J, Konermann S, Severinov K, Zhang F, Koonin EV. Discovery and Functional Characterization of Diverse Class 2 CRISPR-Cas Systems. Mol Cell. 2015 Nov 5;60(3):385-97. doi: 10.1016/j.molcel.2015.10.008. Epub 2015, Oct 22. PMID:26593719 doi:http://dx.doi.org/10.1016/j.molcel.2015.10.008
- ↑ 13.0 13.1 13.2 13.3 13.4 13.5 Jiang F, Zhou K, Ma L, Gressel S, Doudna JA. STRUCTURAL BIOLOGY. A Cas9-guide RNA complex preorganized for target DNA recognition. Science. 2015 Jun 26;348(6242):1477-81. doi: 10.1126/science.aab1452. PMID:26113724 doi:http://dx.doi.org/10.1126/science.aab1452
- ↑ 14.0 14.1 14.2 14.3 14.4 14.5 14.6 14.7 Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012 Aug 17;337(6096):816-21. doi: 10.1126/science.1225829. Epub 2012, Jun 28. PMID:22745249 doi:http://dx.doi.org/10.1126/science.1225829
- ↑ 15.0 15.1 15.2 15.3 Gasiunas G, Barrangou R, Horvath P, Siksnys V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A. 2012 Sep 25;109(39):E2579-86. Epub 2012 Sep 4. PMID:22949671 doi:http://dx.doi.org/10.1073/pnas.1208507109
- ↑ 16.0 16.1 16.2 16.3 16.4 Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013 Feb 28;152(5):1173-83. doi: 10.1016/j.cell.2013.02.022. PMID:23452860 doi:http://dx.doi.org/10.1016/j.cell.2013.02.022
- ↑ 17.0 17.1 17.2 17.3 Bikard D, Jiang W, Samai P, Hochschild A, Zhang F, Marraffini LA. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 2013 Aug;41(15):7429-37. doi: 10.1093/nar/gkt520. Epub 2013, Jun 12. PMID:23761437 doi:http://dx.doi.org/10.1093/nar/gkt520
- ↑ 18.0 18.1 Kuscu C, Arslan S, Singh R, Thorpe J, Adli M. Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nat Biotechnol. 2014 Jul;32(7):677-83. doi: 10.1038/nbt.2916. Epub 2014 May 18. PMID:24837660 doi:http://dx.doi.org/10.1038/nbt.2916
- ↑ Wang R, Zheng H, Preamplume G, Shao Y, Li H. The impact of CRISPR repeat sequence on structures of a Cas6 protein-RNA complex. Protein Sci. 2012 Mar;21(3):405-17. doi: 10.1002/pro.2028. Epub 2012 Feb 9. PMID:22238224 doi:http://dx.doi.org/10.1002/pro.2028
- ↑ Shao Y, Li H. Recognition and Cleavage of a Nonstructured CRISPR RNA by Its Processing Endoribonuclease Cas6. Structure. 2013 Feb 27. pii: S0969-2126(13)00017-8. doi:, 10.1016/j.str.2013.01.010. PMID:23454186 doi:http://dx.doi.org/10.1016/j.str.2013.01.010
- ↑ Reeks J, Sokolowski RD, Graham S, Liu H, Naismith JH, White MF. Structure of a dimeric crenarchaeal Cas6 enzyme with an atypical active site for CRISPR RNA processing. Biochem J. 2013 Mar 25. PMID:23527601 doi:10.1042/BJ20130269
- ↑ Niewoehner O, Jinek M, Doudna JA. Evolution of CRISPR RNA recognition and processing by Cas6 endonucleases. Nucleic Acids Res. 2013 Oct 22. PMID:24150936 doi:http://dx.doi.org/10.1093/nar/gkt922
- ↑ Mulepati S, Heroux A, Bailey S. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science. 2014 Aug 14. pii: 1256996. PMID:25123481 doi:http://dx.doi.org/10.1126/science.1256996
- ↑ Hayes RP, Xiao Y, Ding F, van Erp PB, Rajashankar K, Bailey S, Wiedenheft B, Ke A. Structural basis for promiscuous PAM recognition in type I-E Cascade from E. coli. Nature. 2016 Feb 25;530(7591):499-503. doi: 10.1038/nature16995. Epub 2016 Feb, 10. PMID:26863189 doi:http://dx.doi.org/10.1038/nature16995
- ↑ Marraffini LA. CRISPR-Cas immunity in prokaryotes. Nature. 2015 Oct 1;526(7571):55-61. doi: 10.1038/nature15386. PMID:26432244 doi:http://dx.doi.org/10.1038/nature15386
- ↑ Jinek M, Jiang F, Taylor DW, Sternberg SH, Kaya E, Ma E, Anders C, Hauer M, Zhou K, Lin S, Kaplan M, Iavarone AT, Charpentier E, Nogales E, Doudna JA. Structures of Cas9 Endonucleases Reveal RNA-Mediated Conformational Activation. Science. 2014 Feb 6. PMID:24505130 doi:http://dx.doi.org/10.1126/science.1247997
- ↑ Nishimasu H, Ran FA, Hsu PD, Konermann S, Shehata SI, Dohmae N, Ishitani R, Zhang F, Nureki O. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell. 2014 Feb 27;156(5):935-49. doi: 10.1016/j.cell.2014.02.001. Epub 2014 Feb, 13. PMID:24529477 doi:http://dx.doi.org/10.1016/j.cell.2014.02.001
- ↑ Jiang F, Taylor DW, Chen JS, Kornfeld JE, Zhou K, Thompson AJ, Nogales E, Doudna JA. Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science. 2016 Jan 14. pii: aad8282. PMID:26841432 doi:http://dx.doi.org/10.1126/science.aad8282
- ↑ 29.0 29.1 Nielsen AA, Voigt CA. Multi-input CRISPR/Cas genetic circuits that interface host regulatory networks. Mol Syst Biol. 2014 Nov 24;10:763. doi: 10.15252/msb.20145735. PMID:25422271
- ↑ 30.0 30.1 Didovyk A, Borek B, Hasty J, Tsimring L. Orthogonal Modular Gene Repression in Escherichia coli Using Engineered CRISPR/Cas9. ACS Synth Biol. 2016 Jan 15;5(1):81-8. doi: 10.1021/acssynbio.5b00147. Epub 2015 , Sep 30. PMID:26390083 doi:http://dx.doi.org/10.1021/acssynbio.5b00147
- ↑ 31.0 31.1 Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, Stern-Ginossar N, Brandman O, Whitehead EH, Doudna JA, Lim WA, Weissman JS, Qi LS. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 2013 Jul 18;154(2):442-51. doi: 10.1016/j.cell.2013.06.044. Epub 2013 Jul, 11. PMID:23849981 doi:http://dx.doi.org/10.1016/j.cell.2013.06.044
- ↑ 32.0 32.1 Farzadfard F, Perli SD, Lu TK. Tunable and multifunctional eukaryotic transcription factors based on CRISPR/Cas. ACS Synth Biol. 2013 Oct 18;2(10):604-13. doi: 10.1021/sb400081r. Epub 2013 Sep, 11. PMID:23977949 doi:http://dx.doi.org/10.1021/sb400081r
- ↑ 33.0 33.1 Kiani S, Beal J, Ebrahimkhani MR, Huh J, Hall RN, Xie Z, Li Y, Weiss R. CRISPR transcriptional repression devices and layered circuits in mammalian cells. Nat Methods. 2014 Jul;11(7):723-6. doi: 10.1038/nmeth.2969. Epub 2014 May 5. PMID:24797424 doi:http://dx.doi.org/10.1038/nmeth.2969
![Fig. 1 Overview of the CRISPR-Cas systems. (A) Architecture of class 1 (multiprotein effector complexes) and class 2 (single-protein effector complexes) CRISPR-Cas systems. (B) CRISPR-Cas adaptive immunity is mediated by CRISPR RNAs (crRNAs) and Cas proteins, which form multicomponent CRISPR ribonucleoprotein (crRNP) complexes. The first stage is adaptation, which occurs upon entry of an invading mobile genetic element (in this case, a viral genome). Cas1 (blue) and Cas2 (yellow) proteins select and process the invading DNA, and thereafter, a protospacer (orange) is integrated as a new spacer at the leader end of the CRISPR array [repeat sequences (gray) that separate similar-sized, invader-derived spacers (multiple colors)]. During the second stage, expression, the CRISPR locus is transcribed and the pre-crRNA is processed into mature crRNA guides by Cas (e.g., Cas6) or non-Cas proteins (e.g., RNase III). During the final interference stage, the Cas-crRNA complex scans invading DNA for a complementary nucleic acid target, after which the target is degraded by a Cas nuclease. From](/wiki/images/thumb/f/f2/F1r.jpg/450px-F1r.jpg)