This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1qku
From Proteopedia
| Line 7: | Line 7: | ||
|ACTIVITY= | |ACTIVITY= | ||
|GENE= | |GENE= | ||
| + | |DOMAIN= | ||
| + | |RELATEDENTRY= | ||
| + | |RESOURCES=<span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1qku FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1qku OCA], [http://www.ebi.ac.uk/pdbsum/1qku PDBsum], [http://www.rcsb.org/pdb/explore.do?structureId=1qku RCSB]</span> | ||
}} | }} | ||
| Line 14: | Line 17: | ||
==Overview== | ==Overview== | ||
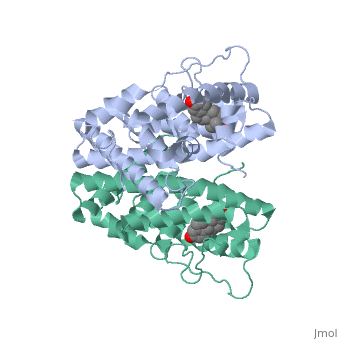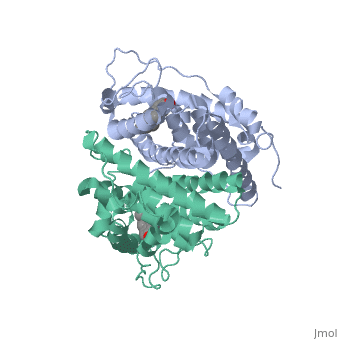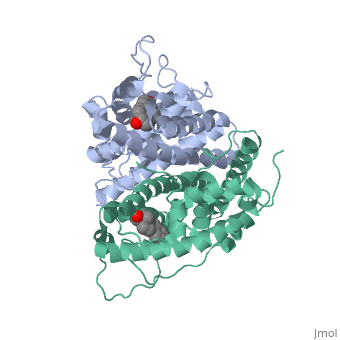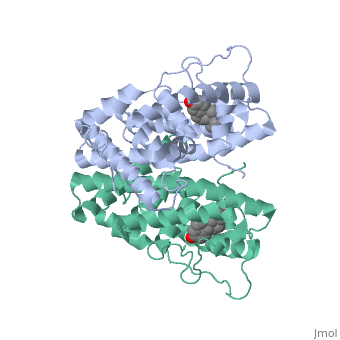
The crystal structure of a triple cysteine to serine mutant ERalpha ligand-binding domain (LBD), complexed with estradiol, shows that despite the presence of a tightly bound agonist ligand, the protein exhibits an antagonist-like conformation, similar to that observed in raloxifen and 4-hydroxytamoxifen-bound structures. This mutated receptor binds estradiol with wild type affinity and displays transcriptional activity upon estradiol stimulation, but with limited potency (about 50%). This partial activity is efficiently repressed in antagonist competition assays. The comparison with available LBD structures reveals key features governing the positioning of helix H12 and highlights the importance of cysteine residues in promoting an active conformation. Furthermore the present study reveals a hydrogen bond network connecting ligand binding to protein trans conformation. These observations support a dynamic view of H12 positioning, where the control of the equilibrium between two stable locations determines the partial agonist character of a given ligand. | The crystal structure of a triple cysteine to serine mutant ERalpha ligand-binding domain (LBD), complexed with estradiol, shows that despite the presence of a tightly bound agonist ligand, the protein exhibits an antagonist-like conformation, similar to that observed in raloxifen and 4-hydroxytamoxifen-bound structures. This mutated receptor binds estradiol with wild type affinity and displays transcriptional activity upon estradiol stimulation, but with limited potency (about 50%). This partial activity is efficiently repressed in antagonist competition assays. The comparison with available LBD structures reveals key features governing the positioning of helix H12 and highlights the importance of cysteine residues in promoting an active conformation. Furthermore the present study reveals a hydrogen bond network connecting ligand binding to protein trans conformation. These observations support a dynamic view of H12 positioning, where the control of the equilibrium between two stable locations determines the partial agonist character of a given ligand. | ||
| - | |||
| - | ==Disease== | ||
| - | Known diseases associated with this structure: Atherosclerosis, susceptibility to OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]], Breast cancer OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]], Estrogen resistance OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]], HDL response to hormone replacement, augmented OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]], Migraine, susceptibility to OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]], Myocardial infarction, susceptibility to OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=133430 133430]] | ||
==About this Structure== | ==About this Structure== | ||
| Line 33: | Line 33: | ||
[[Category: SPINE, Structural Proteomics in Europe.]] | [[Category: SPINE, Structural Proteomics in Europe.]] | ||
[[Category: Wurtz, J M.]] | [[Category: Wurtz, J M.]] | ||
| - | [[Category: EST]] | ||
[[Category: agonism]] | [[Category: agonism]] | ||
[[Category: antagonism]] | [[Category: antagonism]] | ||
| Line 43: | Line 42: | ||
[[Category: structural proteomics in europe]] | [[Category: structural proteomics in europe]] | ||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Sun Mar 30 23:15:07 2008'' |
Revision as of 20:15, 30 March 2008
| |||||||
| , resolution 3.20Å | |||||||
|---|---|---|---|---|---|---|---|
| Sites: | , and | ||||||
| Ligands: | |||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
WILD TYPE ESTROGEN NUCLEAR RECEPTOR LIGAND BINDING DOMAIN COMPLEXED WITH ESTRADIOL
Overview
The crystal structure of a triple cysteine to serine mutant ERalpha ligand-binding domain (LBD), complexed with estradiol, shows that despite the presence of a tightly bound agonist ligand, the protein exhibits an antagonist-like conformation, similar to that observed in raloxifen and 4-hydroxytamoxifen-bound structures. This mutated receptor binds estradiol with wild type affinity and displays transcriptional activity upon estradiol stimulation, but with limited potency (about 50%). This partial activity is efficiently repressed in antagonist competition assays. The comparison with available LBD structures reveals key features governing the positioning of helix H12 and highlights the importance of cysteine residues in promoting an active conformation. Furthermore the present study reveals a hydrogen bond network connecting ligand binding to protein trans conformation. These observations support a dynamic view of H12 positioning, where the control of the equilibrium between two stable locations determines the partial agonist character of a given ligand.
About this Structure
1QKU is a Single protein structure of sequence from Homo sapiens. The following page contains interesting information on the relation of 1QKU with [Estrogen Receptor]. Full crystallographic information is available from OCA.
Reference
Crystal structure of a mutant hERalpha ligand-binding domain reveals key structural features for the mechanism of partial agonism., Gangloff M, Ruff M, Eiler S, Duclaud S, Wurtz JM, Moras D, J Biol Chem. 2001 May 4;276(18):15059-65. Epub 2001 Feb 6. PMID:11278577
Page seeded by OCA on Sun Mar 30 23:15:07 2008
Categories: Estrogen Receptor | Homo sapiens | Single protein | Duclaud, S. | Eiler, S. | Gangloff, M. | Moras, D. | Ruff, M. | SPINE, Structural Proteomics in Europe. | Wurtz, J M. | Agonism | Antagonism | Crystal structure | Estradiol receptor | Spine | Steroid | Structural genomic | Structural proteomics in europe