This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | ==Caspase 6 in ''Homo | + | == '''Caspase-6''' in ''Homo sapiens'' == |
| - | <StructureSection load='4FXO' size='340' side='right' caption=' | + | <StructureSection load='4FXO' size='340' side='right' caption='Caspase-6' scene=''> |
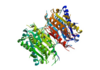
| + | Found at high concentrations in the brain and bordering tissues, Caspase-6 has been implicated in several neurological diseases including Alzheimer's and dementia. It's primarily involved in apoptosis through a largely ambiguous mechanism. It is classified as an [https://en.wikipedia.org/wiki/Endopeptidase]endopeptidase as it cleaves an internal peptide bond of its substrate. It has relatively low specificity in the binding site which allows for a variety of substrates, including other caspase enzymes to bind. Furthermore, it is a part of the cysteine aspartate family, which have these critical amino acid residues in the active site of the enzyme. Caspase-6 has both an inactive zinc-bound conformation and an active ligand-bound conformation, which are largely regulated by variations in zinc concentration. | ||
| + | |||
| + | [[Image:Caspase-6 protein.jpg|100 px|left|thumb|Figure Legend]] | ||
| - | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | ||
[[Image:4FXO.PNG|100 px|left|thumb|This is the figure legend of the thumbnail]] | [[Image:4FXO.PNG|100 px|left|thumb|This is the figure legend of the thumbnail]] | ||
=='''Structure'''== | =='''Structure'''== | ||
| Line 23: | Line 25: | ||
===Self cleavage=== | ===Self cleavage=== | ||
| - | ==''' | + | == Inhibition == |
| + | :'''Zinc Inhibition''' | ||
| + | Primary inhibition of Caspase-6 occurs when a zinc ion binds to the exosite containing Lys-36, Glu-244, and His-287 of the active dimer. In addition to these residues, the zinc interacts with one water molecule from the cytoplasm. It has been proposed that helices of the active dimer must rotate or move in some other way to provide these ideal interactions with zinc. This subtle shift is most likely the cause for allosteric inhibition. As the helices move to bind zinc, the amino acids of the active site become misaligned. The altered positions of the amino acids no longer provide ideal interactions for incoming substrates. After zinc binds, no new substrates enter the active site. Thus, Caspase-6 is effectively inhibited. | ||
| + | |||
| + | :'''Phosphorylation''' | ||
| + | The function of Caspase-6 can be inhibited by phosphorylation of Ser-257. The exact mechanism of this reaction remains unidentified at the time of publication, but proceeds when ARK5 kinase is present. This modification can occur before and after zymogen activation or auto-processing. The phosphoryl group inhibits Caspase-6 through steric interference. When Ser-257 is phosphorylated, the amino acid residue interacts with Pro-201, causing a shift in the helices of Caspase-6. The shift misaligns and disrupts residues found in the active site. This conformational difference prevents the inter-subunit loop from entering during zymogen activation and the self-cleaved active dimer cannot be formed. Additionally, no new substrate is able to enter the active site. | ||
| + | |||
| + | :'''Zymogen Activation''' | ||
===Zinc Inhibition=== | ===Zinc Inhibition=== | ||
Revision as of 01:22, 2 April 2017
Caspase-6 in Homo sapiens
| |||||||||||
References
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966951/ (self cleavage article)
http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien)