This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
| Line 13: | Line 13: | ||
=='''Activation of Caspase-6'''== | =='''Activation of Caspase-6'''== | ||
| - | Caspase-6 has a small pro-domain. It shares 41% sequence identity with Caspase-3 and 37% sequence identity with Caspase-7 [[https://www.ncbi.nlm.nih.gov/pubmed/25340928]. Both of these caspases are classified as effectors and because of it's similarites to these other Caspases, Caspase-6 is also classified as an effector. Caspase-6, does however, have many unique features compared to the other effectors, it has similar substrate specificity to that of initiator Caspases-8 and -9. Inhibitors of Apoptosis or IAPs, which are known to inhibit Caspase-3, -7, and -9, do not inhibit Caspase-6. Caspase-6 is known to undergo self-processing and activation in vitro and in vivo. High activity of Caspase-6 protein do not induce apoptosis of in HEK293 cells. Caspase-6 also has a relatively low zymogenicty, which is the ratio of activity for cleaved protein to the activity of uncleaved protein, of about 200, which is comparable to the zymogenicities of Caspase-8 and Caspase-9, which are both classified as initiator caspases. Caspase-6’s zymogenicity is also much lower than Caspase-3, which is another effector. This is interesting because the Caspase-6 protein shows low activity when it is not cleaved, similar to the initiator caspases, but Caspase-3, another effector, has basically no zymogen activity. Caspase-6 is classified as an effector, but it can also act as an initiator and cleave Caspases-2 and -8. It can also induce the mitochondrial membrane to become permeable, which leads to cytochrome c release and activation of other effector caspases. | ||
===Structural Units involved in autoactivation=== | ===Structural Units involved in autoactivation=== | ||
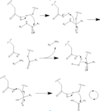
| - | + | Before Caspase-6 is a functional and active dimer, the enzyme exists as a procaspase. This precursor enzyme is modified by self-processing, a characteristic unique to Caspase-6. The unprocessed enzyme contains a small and large subunit, a pro domain, as well as a intersubunit linker. To become active, the intersubunit linker binds to the active site, where it is then cleaved. Other cleavages must occur as well for the enzyme to become active, specifically at TETD23 of the pro-domain*, DVVD179, and TEVD193 amino acid sequences. The pro-domain is released after the cleavage at TETD23, which then allows the two subunits to interact to form the active dimer. | |
| + | Caspase-6 can also be activated by other caspases as an alternate to auto-activation. | ||
| + | |||
| + | *Cleavage at these sites occurs in a specific sequence. To begin, the site within the pro-domain, TETD23, must be cleaved. This cleavage is then followed by either DVVD179 or TEVD193. It has been proposed that thus sequence of cleavage is due to the structure of Caspase-6, which allows the pro-domain to be more available to enter the active site. To some extent, the pro-domain inhibits Caspase-6's ability to cleave the intersubunit loop and self-activate, but this also happens in a currently unknown mechanism. | ||
===Caspase-6 Self-Cleavage Autoactivation=== | ===Caspase-6 Self-Cleavage Autoactivation=== | ||
| - | It has been found that Caspase-6 can undergo activation without any other caspases | + | It has been found that Caspase-6 can undergo activation without any other caspases, so there is a suggested self-cleavage mechanism for Caspase-6. The intramolecular cleavage of TEVD193 is essential for the initiation caspase-6 activation without Caspase-3 present. The prodomain somehow inhibits the intramolecular cleavage of TEVD193, but currently the mechanism is not known. The TETD23 and TEVD193 cleavage sites are similar, but the TETD23 cleavage site is always cleaved before TEVD193. This indicates that the TETD23 cleavage site is always more readily available for cleavage. The result of the TETD23 cleavage site priority is that the prodomain acts as a “suicide protector”, which protects the TEVD193 cleavage site from intermolecular self-cleavage. This protection is useful when there are low levels of protein, such as when it is in vivo, it also helps explain why the prodomain inhibits self-activation in vivo, but not in vitro. |
| Line 28: | Line 30: | ||
===Phosphorylation=== | ===Phosphorylation=== | ||
The function of Caspase-6 can be inhibited by phosphorylation of Ser-257. The exact mechanism of this reaction remains unidentified at the time of publication, but proceeds when ARK5 kinase is present. This modification can occur before and after zymogen activation or auto-processing. The phosphoryl group inhibits Caspase-6 through steric interference. When Ser-257 is phosphorylated, the amino acid residue interacts with Pro-201, causing a shift in the helices of Caspase-6. The shift misaligns and disrupts residues found in the active site. This conformational difference prevents the inter-subunit loop from entering during zymogen activation and the self-cleaved active dimer cannot be formed. Additionally, no new substrate is able to enter the active site. | The function of Caspase-6 can be inhibited by phosphorylation of Ser-257. The exact mechanism of this reaction remains unidentified at the time of publication, but proceeds when ARK5 kinase is present. This modification can occur before and after zymogen activation or auto-processing. The phosphoryl group inhibits Caspase-6 through steric interference. When Ser-257 is phosphorylated, the amino acid residue interacts with Pro-201, causing a shift in the helices of Caspase-6. The shift misaligns and disrupts residues found in the active site. This conformational difference prevents the inter-subunit loop from entering during zymogen activation and the self-cleaved active dimer cannot be formed. Additionally, no new substrate is able to enter the active site. | ||
| - | |||
| - | ===Zymogen Activation=== | ||
| Line 53: | Line 53: | ||
== References == | == References == | ||
<references/> | <references/> | ||
| - | + | Wang, Xiao-Jun, Qin Cao, Yan Zhang, and Xiao-Dong Su. "Activation and Regulation of Caspase-6 and Its Role in Neurodegenerative Diseases." Annual Review of Pharmacology and Toxicology 55.1 (2015): 553-72. Web. | |
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966951/ (self cleavage article) | https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966951/ (self cleavage article) | ||
http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien) | http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien) | ||
Revision as of 21:49, 2 April 2017
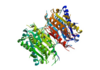
Caspase-6 in Homo sapiens
| |||||||||||
References
Wang, Xiao-Jun, Qin Cao, Yan Zhang, and Xiao-Dong Su. "Activation and Regulation of Caspase-6 and Its Role in Neurodegenerative Diseases." Annual Review of Pharmacology and Toxicology 55.1 (2015): 553-72. Web.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966951/ (self cleavage article)
http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien)