This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
| Line 14: | Line 14: | ||
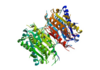
In order to function as an endoprotease, Caspase-6 binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and tubulins [https://en.wikipedia.org/wiki/Tubulin], in its active site.[[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding grove in Caspase-6. Blue - catalytic residues | In order to function as an endoprotease, Caspase-6 binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and tubulins [https://en.wikipedia.org/wiki/Tubulin], in its active site.[[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding grove in Caspase-6. Blue - catalytic residues | ||
yellow - ligand | yellow - ligand | ||
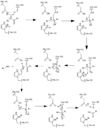
| - | red - generic surface]] This binding groove contains three critical amino acid residues necessary to perform cleavage of the peptide bonds. Together,<scene name='75/752344/His121_real/1'>His-121</scene>, <scene name='75/752344/Glu123_real/1'>Glu-123</scene>, and <scene name='75/752344/Cys163_real/1'>Cys-163</scene> form a <scene name='75/752344/Catalytic_triad_real/1'>catalytic triad</scene>[[Image:The real caspase mechanism.jpg|100 px|right|thumb|active site mechanism]]. In the theorized mechanism, His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. | + | red - generic surface]] This binding groove contains three critical amino acid residues necessary to perform cleavage of the peptide bonds. Together, <scene name='75/752344/His121_real/1'>His-121</scene>, <scene name='75/752344/Glu123_real/1'>Glu-123</scene>, and <scene name='75/752344/Cys163_real/1'>Cys-163</scene> form a <scene name='75/752344/Catalytic_triad_real/1'>catalytic triad</scene>[[Image:The real caspase mechanism.jpg|100 px|right|thumb|active site mechanism]]. In the theorized mechanism, His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. |
===Zinc Exosite=== | ===Zinc Exosite=== | ||
| - | Caspase-6 function is inhibited by the binding of a <scene name='75/752344/Zinc_caspase-6/1'> | + | Caspase-6 function is inhibited by the binding of a <scene name='75/752344/Zinc_caspase-6/1'>zinc</scene> ion[https://en.wikipedia.org/wiki/Zinc], which binds to an <scene name='75/752344/Caspase6_allosteric_site/1'>allosteric site</scene> instead of the <scene name='75/752344/Caspase6_allostericactiv_site/1'>active site</scene>. This allosteric site is located on the outside of the protein and is distal to the active site. The zinc ion is bound to <scene name='75/752344/Caspase6_allosteric_site_resid/1'>three amino acid residues</scene>, Lys-36, Glu-244, and His-287. Once the ion is bound to the protein, it is then stabilized by a <scene name='75/752344/H20_zinc_binding_casp/1'>water molecule</scene>. The binding of zinc at the exosite is suggested to cause a conformational change in the protein from an <scene name='75/752344/Catalytic_triad_real/1'>active state</scene> to an <scene name='75/752344/Inactive_catalytic_triad_casp/1'>inactive state</scene> that misaligns catalytic residues and inhibits activity of the enzyme. The residues in the active site no longer provide ideal interactions with the substrate and therefore, substrate does not bind. Zinc binding to the exosite is tightly regulated as it inhibits Caspase-6's critical role in initiation of apoptosis. |
=='''Activation of Caspase-6'''== | =='''Activation of Caspase-6'''== | ||
Revision as of 00:40, 4 April 2017
Caspase-6 in Homo sapiens
| |||||||||||
References
- ↑ Wang XJ, Cao Q, Zhang Y, Su XD. Activation and regulation of caspase-6 and its role in neurodegenerative diseases. Annu Rev Pharmacol Toxicol. 2015;55:553-72. doi:, 10.1146/annurev-pharmtox-010814-124414. Epub 2014 Oct 17. PMID:25340928 doi:http://dx.doi.org/10.1146/annurev-pharmtox-010814-124414
- ↑ 2.0 2.1 Velazquez-Delgado EM, Hardy JA. Zinc-Mediated Allosteric Inhibition of Caspase-6. J Biol Chem. 2012 Aug 13. PMID:22891250 doi:http://dx.doi.org/10.1074/jbc.M112.397752
- ↑ Velazquez-Delgado EM, Hardy JA. Phosphorylation regulates assembly of the caspase-6 substrate-binding groove. Structure. 2012 Apr 4;20(4):742-51. Epub 2012 Apr 3. PMID:22483120 doi:10.1016/j.str.2012.02.003
Wang, Xiao-Jun, Qin Cao, Yan Zhang, and Xiao-Dong Su. "Activation and Regulation of Caspase-6 and Its Role in Neurodegenerative Diseases." Annual Review of Pharmacology and Toxicology 55.1 (2015): 553-72. Web.
Wang XJ, Cao Q, Liu X, Wang KT, Mi W, et al. 2010. Crystal structures of human caspase 6 reveal a new mechanism for intramolecular cleavage self-activation. EMBO Rep. 11: 841–47
(self cleavage article)
http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien)